Hereditas(Beijing) ›› 2020, Vol. 42 ›› Issue (8): 752-759.doi: 10.16288/j.yczz.19-272
• Review • Previous Articles Next Articles
Progress on phage genomics of Pseudomonas spp.
Zhiwei Xu, Yunlin Wei, Xiuling Ji( )
)
- Faculty of Life Science and Technology, Kunming University of Science and Technology, Kunming 650500, China
-
Received:2020-04-26Revised:2020-07-10Online:2020-08-20Published:2020-07-29 -
Contact:Ji Xiuling E-mail:jixiuling1023@126.com -
Supported by:Supported by the National Natural Science Foundation of China Nos(31860147);Supported by the National Natural Science Foundation of China Nos(31700324)
Cite this article
Zhiwei Xu, Yunlin Wei, Xiuling Ji. Progress on phage genomics of Pseudomonas spp.[J]. Hereditas(Beijing), 2020, 42(8): 752-759.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
| [1] |
Silby MW, Winstanley C, Godfrey SA, Levy SB, Jackson RW . Pseudomonas genomes: diverse and adaptable. FEMS Microbiol Rev, 2011,35(4):652-680.
doi: 10.1111/j.1574-6976.2011.00269.x pmid: 21361996 |
| [2] | Czajkowski R, Ozymko Z, de Jager V, Siwinska J, Smolarska A, Ossowicki A, Narajczyk M, Lojkowska E. Genomic, proteomic and morphological characterization of two novel broad host lytic bacteriophages ΦPD10.3 and ΦPD23.1 infecting pectinolytic Pectobacterium spp. and Dickeya spp. PLoS ONE, 2015,10(3):1-23. |
| [3] |
Olszak T, Latka A, Roszniowski B, Valvano MA, Drulis- Kawa Z . Phage life cycles behind bacterial biodiversity. Curr Med Chem, 2017,24(36):3987-4001.
doi: 10.2174/0929867324666170413100136 pmid: 28412903 |
| [4] |
Alseth EO, Pursey E, Luján AM, McLeod I, Rollie C, Westra ER. Bacterial biodiversity drives the evolution of CRISPR-based phage resistance. Nature, 2019,574(7779):549-552.
doi: 10.1038/s41586-019-1662-9 pmid: 31645729 |
| [5] | Li TM, Du B . CRISPR-Cas system and coevolution of bacteria and phages. Hereditas(Beijing), 2011,33(3):213-218. |
| 李铁民, 杜波 . CRISPR-Cas系统与细菌和噬菌体的共进化. 遗传, 2011,33(3):213-218. | |
| [6] |
Ceyssens PJ, Hertveldt K, Ackermann HW, Noben JP, Demeke M, Volckaert G, Lavigne R . The intron-containing genome of the lytic Pseudomonas phage LUZ24 resembles the temperate phage PaP3. Virology, 2008,377(2):233-238.
doi: 10.1016/j.virol.2008.04.038 pmid: 18519145 |
| [7] |
Ceyssens PJ, Mesyanzhinov V, Sykilinda N, Briers Y, Roucourt B, Lavigne R, Robben J, Domashin A, Miroshnikov K, Volckaert G, Hertveldt K . The genome and structural proteome of YuA, a new Pseudomonas aeruginosa phage resembling M6. Journal of bacteriology, 2007,190(4):1429-1435.
doi: 10.1128/JB.01441-07 pmid: 18065532 |
| [8] |
Hatfull GF, Pedulla ML, Jacobs-Sera D, Cichon PM, Foley A, Ford ME, Gonda RM, Houtz JM, Hryckowian AJ, Kelchner VA, Namburi S, Pajcini KV, Popovich MG, Schleicher DT, Simanek BZ, Smith AL, Zdanowicz GM, Kumar V, Peebles CL, Jacobs WR Jr, Lawrence JG, Hendrix RW . Exploring the mycobacteriophage metaproteome: phage genomics as an educational platform. PLoS Genet, 2006,2(6):e92.
doi: 10.1371/journal.pgen.0020092 pmid: 16789831 |
| [9] |
Amgarten D, Martins LF, Lombardi KC, Antunes LP, de Souza APS, Nicastro GG, Kitajima EW, Quaggio RB, Upton C, Setubal JC, da Silva AM. Three novel Pseudomonas phages isolated from composting provide insights into the evolution and diversity of tailed phages. BMC Genomics, 2017,18(1):346-363.
doi: 10.1186/s12864-017-3729-z pmid: 28472930 |
| [10] |
Wang PW, Chu LD, Guttman DS . Complete sequence and evolutionary genomic analysis of the pseudomonas aeruginosa transposable bacteriophage D3112. J Bacteriol, 2004,186(2):400-410.
doi: 10.1128/jb.186.2.400-410.2004 pmid: 14702309 |
| [11] |
Ha AD, Denver DR . Comparative genomic analysis of 130 bacteriophages Infecting bacteria in the genus Pseudomonas. Front Microbiol, 2018,9:1456.
doi: 10.3389/fmicb.2018.01456 pmid: 30022972 |
| [12] |
Wittmann J, Dreiseikelmann B, Rohde M, Meier-Kolthoff JP, Bunk B, Rohde C . First genome sequences of achromobacter phages reveal new members of the N4 family. Virol J, 2014,11(1):2-15.
doi: 10.1186/1743-422X-11-2 |
| [13] |
Sillankorva S, Kluskens LD, Lingohr EJ, Kropinski AM, Neubauer P, Azeredo J . Complete genome sequence of the lytic Pseudomonas fluorescens phage ϕIBB-PF7A. Virol J, 2011,8:142.
doi: 10.1186/1743-422X-8-142 pmid: 21439081 |
| [14] |
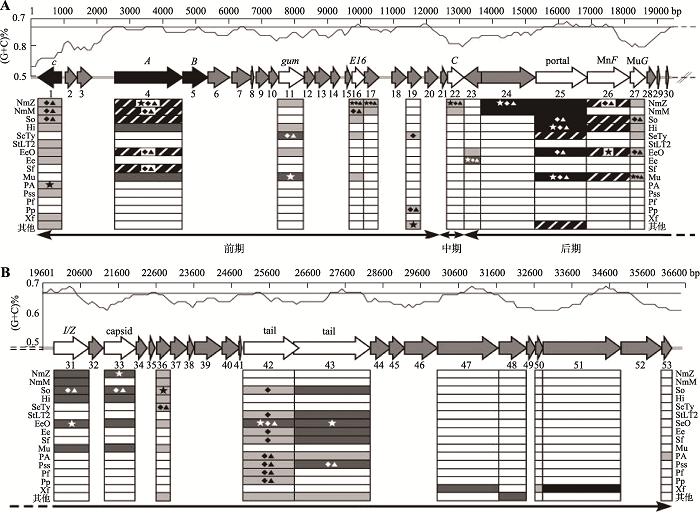
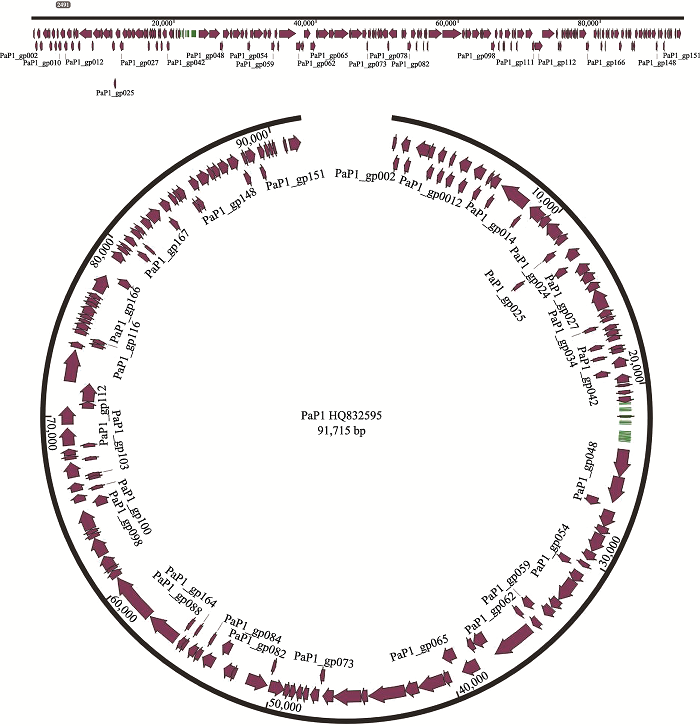
Lu SG, Le S, Tan YL, Zhu JM, Li M, Rao XC, Zou LY, Li S, Wang J, Jin XL, Huang GT, Zhang L, Zhao X, Hu FQ . Genomic and proteomic analyses of the terminally redundant genome of the Pseudomonas aeruginosa phage PaP1: establishment of genus PaP1-like phages. PLoS One, 2013,8(5):e62933.
doi: 10.1371/journal.pone.0062933 pmid: 23675441 |
| [15] |
Kwan T, Liu J, Dubow M, Gros P, Pelletier J . Comparative genomic analysis of 18 Pseudomonas aeruginosa bacteriophages. J Bacteriol, 2006,188(3):1184-1187.
doi: 10.1128/JB.188.3.1184-1187.2006 pmid: 16428425 |
| [16] |
Watkins SC, Sible E, Putonti C . Pseudomonas PB1-Like phages: whole genomes from metagenomes offer insight into an abundant group of bacteriophages. Viruses, 2018,10(1):331-344.
doi: 10.3390/v10060331 |
| [17] |
Li M, Chen XR, Ma YS, Li ZB, Zhao QC . Complete genome sequence of PFP1, a novel T7-like Pseudomonas fluorescens bacteriophage. Arch Virol, 2018,163(1):3423-3426.
doi: 10.1007/s00705-018-3979-3 |
| [18] |
Lu SG, Le S, Tan YL, Li M, Liu C, Zhang KB, Huang JJ, Chen HM, Rao XC, Zhu JM, Zou LY, Ni QS, Li S, Wang J, Jin XL, Hu QW, Yao XY, Zhao X, Zhang L, Huang GT, Hu FQ . Unlocking the mystery of the hard-to-sequence phage genome: PaP1 methylome and bacterial immunity. BMC Genomics, 2014,15(1):803-815.
doi: 10.1186/1471-2164-15-803 |
| [19] |
Aparicio T, de Lorenzo V, Martínez-García E . Improved thermotolerance of genome-reduced Pseudomonas putida EM42 enables effective functioning of the PL/cI857 system. Biotechnol J, 2019,14(1):1800483.
doi: 10.1002/biot.v14.1 |
| [20] |
Zhang FJ, Huang KC, Yang XJ, Sun L, You JJ, Pan XW, Cui XL, Yang HJ . Characterization of a novel lytic podovirus O4 of Pseudomonas aeruginosa. Arch Virol, 2018,163(1):2377-2383.
doi: 10.1007/s00705-018-3866-y |
| [21] | Carson S, Bruff E, DeFoor W, Dums J, Groth A, Hatfield T, Iyer A, Joshi K, McAdams S, Miles D, Miller D, Oufkir A, Raynor B, Riley S, Roland S, Rozier H, Talley S, Miller ES. Genome sequences of six paenibacillus larvae Siphoviridae Phages. Genome Announc, 2015,3(3):101-115. |
| [22] |
Tang CF, Deng CJ, Zhang Y, Xiao C, Wang J, Rao XC, Hu FQ, Lu SG . Characterization and genomic analyses of Pseudomonas aeruginosa podovirus TC6: establishment of genus Pa11virus. Front Microbiol, 2018,9:2561.
doi: 10.3389/fmicb.2018.02561 pmid: 30410478 |
| [23] | Xu B, GAO J, GUO XK, Qin JH . Study on the biological and genomic characteristics of Pseudomonas aeruginosa phage D204. J Shanghai Jiaotong Univ, 2016,36(1):1-5. |
| 徐彬, 高晶, 郭晓奎, 秦金红 . 铜绿假单胞菌噬菌体D204的生物学特性和基因组学研究. 上海交通大学学报, 2016,36(1):1-5. | |
| [24] | Cui XL . Screening of genes related to pseudomonas aeruginosa response to multiple strains of phage infection and functional annotation of phage C11 genome [Dissertation]. Sc i Tech of Tianjin Univ, 2016. |
| 崔晓莉 . 铜绿假单胞菌应答多株噬菌体感染相关基因的筛选及噬菌体C11基因组的功能注释[学位论文]. 天津科技大学, 2016. | |
| [25] |
Sharma S, Chatterjee S, Datta S, Prasad R, Dubey D, Prasad RK, Vairale MG . Bacteriophages and its applications: an overview. Folia Microbiol (Praha), 2017,62(1):17-55.
doi: 10.1007/s12223-016-0471-x |
| [26] |
Cao HL, Lai Y, Bougouffa S, Xu ZL, Yan AX . Comparative genome and transcriptome analysis reveals distinctive surface characteristics and unique physiological potentials of Pseudomonas aeruginosa ATCC 27853. BMC Genomics, 2017,18(1):459-475.
doi: 10.1186/s12864-017-3842-z pmid: 28606056 |
| [27] |
De Smet J, Hendrix H, Blasdel BG, Danis-Wlodarczyk K, Lavigne R . Pseudomonas predators: understanding and exploiting phage-host interactions. Nat Rev Microbiol, 2017,15(9):517-530.
doi: 10.1038/nrmicro.2017.61 pmid: 28649138 |
| [28] | Liang CJ, Meng FM, Ai YC . CRISPR/Cas systems in genome engineering of bacteriophages. Hereditas(Beijing), 2018,40(5):378-389. |
| 梁彩娇, 孟繁梅, 艾云灿 . 基于CRISPR/Cas系统的噬菌体基因组编辑. 遗传, 2018,40(5):378-389. | |
| [29] |
Pawluk A, Bondy-Denomy J, Cheung VH, Maxwell KL, Davidson AR . A new group of phage anti-CRISPR genes inhibits the type I-E CRISPR-Cas system of Pseudomonas aeruginosa. mBio, 2014,5(2):e00896.
doi: 10.1128/mBio.00896-14 pmid: 24736222 |
| [30] |
Habusha M, Tzipilevich E, Fiyaksel O, Ben-Yehuda S . A mutant bacteriophage evolved to infect resistant bacteria gained a broader host range. Mol Microbiol, 2019,111(6):1463-1475.
doi: 10.1111/mmi.14231 pmid: 30811056 |
| [31] |
Holguín AV, Cárdenas P, Prada-Peñaranda C, Rabelo Leite L, Buitrago C, Clavijo V, Oliveira G, Leekitcharoenphon P, Møller Aarestrup F, Vives MJ . Host resistance, genomics and population dynamics in a salmonella Eenteritidis and phage system. Viruses, 2019,11(2):188.
doi: 10.3390/v11020188 |
| [32] | Li M, Cheng FY, Gong LY, Xiang H . Systematic discovery of novel prokaryotic defense systems: progress and prospects. Hereditas(Beijing), 2018,40(4):259-265. |
| 李明, 程飞跃, 龚路遥, 向华 . 微生物新型防御系统的系统性发现与展望. 遗传, 2018,40(4):259-265. | |
| [33] | Gao SH, Yu HY, Wu SY, Wang S, Geng JN, Luo YF, Hu SN . Advances of sequencing and assembling technologies for complex genomes. Hereditas(Beijing), 2018,40(11):944-963. |
| 高胜寒, 禹海英, 吴双阳, 王森, 耿佳宁, 骆迎峰, 胡松年 . 复杂基因组测序技术研究进展. 遗传, 2018,40(11):944-963. | |
| [34] |
Sternberg SH, Richter H, Charpentier E, Qimron U . Adaptation in CRISPR-Cas systems. Mol Cell, 2016,61(6):797-808.
doi: 10.1016/j.molcel.2016.01.030 pmid: 26949040 |
| [1] | Zhong Bian, Dongping Cao, Wenshu Zhuang, Shuwei Zhang, Qiaoquan Liu, Lin Zhang. Revelation of rice molecular design breeding: the blend of tradition and modernity [J]. Hereditas(Beijing), 2023, 45(9): 718-740. |
| [2] | Xiaoping Lian, Guangfu Huang, Yujiao Zhang, Jing Zhang, Fengyi Hu, Shilai Zhang. The discovery and utilization of favorable genes in Oryza longistaminata [J]. Hereditas(Beijing), 2023, 45(9): 765-780. |
| [3] | Jun Ma, Anping Fan, Wusheng Wang, Jinchuan Zhang, Xiaojun Jiang, Ruijun Ma, Sheqiang Jia, Fei Liu, Chuchao Lei, Yongzhen Huang. Analysis of genetic diversity and genetic structure of Qinchuan cattle conservation population using whole-genome resequencing [J]. Hereditas(Beijing), 2023, 45(7): 602-616. |
| [4] | Dantong Xu, Yifei Wang, Jiali Cai, Wentao Gong, Xiangchun Pan, Yuhan Tian, Qingpeng Shen, Jiaqi Li, Xiaolong Yuan. Study on detection of CNVs using human whole genome bisulfite sequencing data [J]. Hereditas(Beijing), 2023, 45(4): 324-340. |
| [5] | Zhongsheng Wu, Yu Gao, Yongtao Du, Song Dang, Kangmin He. The protocol of tagging endogenous proteins with fluorescent tags using CRISPR-Cas9 genome editing [J]. Hereditas(Beijing), 2023, 45(2): 165-175. |
| [6] | Meizhen Liu, Liren Wang, Yongmei Li, Xueyun Ma, Honghui Han, Dali Li. Generation of genetically modified rat models via the CRISPR/Cas9 technology [J]. Hereditas(Beijing), 2023, 45(1): 78-87. |
| [7] | Xiuli Chen, Haiyan Huang, Qiang Wu. Targeted deletion of 5′HS2 enhancer of β-globin locus control region in K562 cells [J]. Hereditas(Beijing), 2022, 44(9): 783-797. |
| [8] | Mengxuan Xu, Ming Zhou. Advances of RNA polymerase IV in controlling DNA methylation and development in plants [J]. Hereditas(Beijing), 2022, 44(7): 567-580. |
| [9] | Wanjing Ping, Yichen Liu, Qiaomei Fu. Exploring the evolution of archaic humans through sedimentary ancient DNA [J]. Hereditas(Beijing), 2022, 44(5): 362-369. |
| [10] | Jiming Xu, Jianshu Zhu, Mengzhen Li, Han Hu, Chuanzao Mao. Progress on methods for acquiring flanking genomic sequence [J]. Hereditas(Beijing), 2022, 44(4): 313-321. |
| [11] | Jie Kou, Yan Li, Peng Wang, Hong Liu, Jiawen Liu, Juan Wang, Ye Wang, Liang Zhang, Fujun Shen. Optimization of microsatellite genotyping system used for genetic diversity evaluation of Ailuropoda melanoleuca [J]. Hereditas(Beijing), 2022, 44(3): 253-266. |
| [12] | Cong Liu, Jiani Feng, Weiwei Li, Weiwei Zhu, Yunxin Xue, Dai Wang, Xilin Zhao. Maintenance of dNTP pool homeostasis and genomic stability [J]. Hereditas(Beijing), 2022, 44(2): 96-106. |
| [13] | Cong Zhou, Qiangwei Zhou, Sheng Cheng, Guoliang Li. Research progress of CTCF in mediating 3D genome formation and regulating gene expression [J]. Hereditas(Beijing), 2021, 43(9): 816-821. |
| [14] | Changgui Lei, Xueyuan Jia, Wenjing Sun. Establish six-gene prognostic model for glioblastoma based on multi-omics data of TCGA database [J]. Hereditas(Beijing), 2021, 43(7): 665-679. |
| [15] | Jing Jiang, Anqi Zhao, Tian Xie, Shuwei Chen, Jinsong Li. Construction of genome-wide protein tagging cell and mouse libraries [J]. Hereditas(Beijing), 2021, 43(7): 704-714. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||