Hereditas(Beijing) ›› 2020, Vol. 42 ›› Issue (12): 1168-1177.doi: 10.16288/j.yczz.20-069
• Review • Previous Articles Next Articles
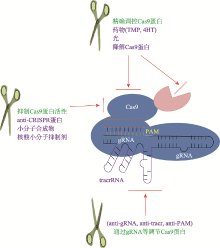
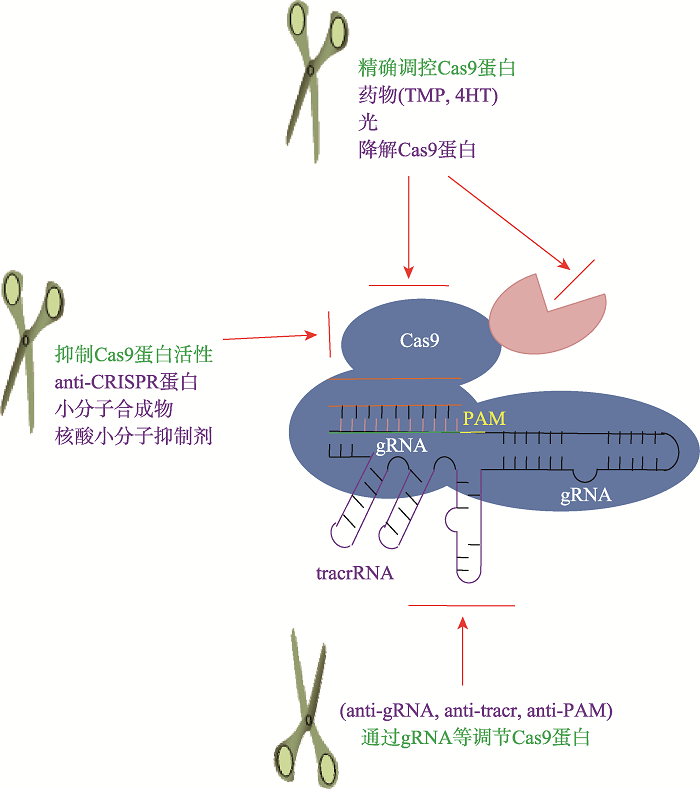
Advances in precise regulation of CRISPR/Cas9 gene editing technology
Junxia Cao1, Youliang Wang2, Zhengxu Wang1,3( )
)
- 1. Biotherapy Center, the Seventh Medical Center of PLA General Hospital, Beijing 100700, China
2. Beijing Institute of Biotechnology, 20 Dongdajie, Beijing 100071, China
3. The Faculty of Hepato-Pancreato-Biliary Surgery, the First Medical Center of Chinese PLA General Hospital, Beijing 100853, China
-
Received:2020-03-13Revised:2020-11-02Online:2020-12-17Published:2020-11-25 -
Contact:Wang Zhengxu E-mail:zhxuwang@qq.com -
Supported by:Supported by the 13th Five Year Research Project No(19LBJ1029C)
Cite this article
Junxia Cao, Youliang Wang, Zhengxu Wang. Advances in precise regulation of CRISPR/Cas9 gene editing technology[J]. Hereditas(Beijing), 2020, 42(12): 1168-1177.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
| [1] | 2013 Runners-Up . Genetic microsurgery for the masses. Science, 2013,342(6165):1434-1435. |
| [2] | Travis J . Breakthrough of the year: CRISPR makes the cut. Science, 2015,350(6267):1456-1457. |
| [3] | Nature Editor . The scientific events that shaped the decade. Nature, 2019,576(7787):337-338. |
| [4] | Enache OM, Rendo V, Abdusamad M, Lam D, Davison D, Pal S, Currimjee N, Hess J, Pantel S, Nag A, Thorner AR, Doench JG, Vazquez F, Beroukhim R, Golub TR, Ben-David U . Cas9 activates the p53 pathway and selects for p53-inactivating mutations. Nat Genet, 2020,52(7):662-668. |
| [5] | Ledford H . Quest to use CRISPR against disease gains ground. Nature, 2020,577(7789):156. |
| [6] | Cyranoski D . What CRISPR-baby prison sentences mean for research. Nature, 2020,577(7789):154-155. |
| [7] | Hwang S, Maxwell KL . Meet the anti-CRISPRs: widespread protein inhibitors of CRISPR-Cas systems. CRISPR J, 2019,2(1):23-30. |
| [8] | Nakamura M, Srinivasan P, Chavez M, Carter MA, Dominguez AA, La Russa M, Lau MB, Abbott TR, Xu XS, Zhao DH, Gao YC, Kipniss NH, Smolke CD, Bondy- Denomy J, Qi LS . Anti-CRISPR-mediated control of gene editing and synthetic circuits in eukaryotic cells. Nat Commun, 2019,10(1):194. |
| [9] | Adli M . The CRISPR tool kit for genome editing and beyond. Nat Commun, 2018,9(1):1911. |
| [10] | Tang LC, Gu F . Next-generation CRISPR-Cas for genome editing: focusing on the Cas protein and PAM. Hereditas (Beijing), 2020, 42(3):236-249. |
| 唐连超, 谷峰 . CRISPR-Cas基因编辑系统升级: 聚焦Cas蛋白和PAM. 遗传, 2020,42(3):236-249. | |
| [11] | Rees HA, Liu DR . Base editing: precision chemistry on the genome and transcriptome of living cells. Nat Rev Genet, 2018,19(12):770-788. |
| [12] | Cyranoski D . CRISPR gene-editing tested in a person for the first time. Nature, 2016,539(7630):479. |
| [13] | Chen YO, Bao Y, Ma HZ, Yi ZY, Zhou Z, Wei WS . Gene editing technology and its research progress in China. Hereditas (Beijing), 2018,40(10):900-915. |
| 陈一欧, 宝颖, 马华峥, 伊宗裔, 周卓, 魏文胜 . 基因编辑技术及其在中国的研究发展. 遗传, 2018,40(10):900-915. | |
| [14] | Xu L, Wang J, Liu YL, Xie LF, Su B, Mou DL, Wang LT, Liu TT, Wang XB, Zhang B, Zhao L, Hu LD, Ning HM, Zhang YF, Deng K, Liu LF, Lu XF, Zhang T, Xu J, Li C, Wu H, Deng HK, Chen H . CRISPR-edited stem cells in a patient with HIV and acute lymphocytic leukemia. N Engl J Med, 2019,381(13):1240-1247. |
| [15] | Wu YX, Zeng J, Roscoe BP, Liu PP, Yao QM, Lazzarotto CR, Clement K, Cole MA, Luk K, Baricordi C, Shen AH, Ren C, Esrick EB, Manis JP, Dorfman DM, Williams DA, Biffi A, Brugnara C, Biasco L, Brendel C, Pinello L, Tsai SQ, Wolfe SA, Bauer DE . Highly efficient therapeutic gene editing of human hematopoietic stem cells. Nat Med, 2019,25(5):776-783. |
| [16] | Qasim W, Zhan H, Samarasinghe S, Adams S, Amrolia P, Stafford S, Butler K, Rivat C, Wright G, Somana K, Ghorashian S, Pinner D, Ahsan G, Gilmour K, Lucchini G, Inglott S, Mifsud W, Chiesa R, Peggs KS, Chan L, Farzeneh F, Thrasher AJ, Vora A, Pule M, Veys P. Molecular remission of infant B-ALL after infusion of universal TALEN gene-edited CAR T cells. Sci Transl Med, 2017, 9(374): eaaj2013. |
| [17] | Cornu TI, Mussolino C, Cathomen T . Refining strategies to translate genome editing to the clinic. Nat Med, 2017,23(4):415-423. |
| [18] | Howard HC, van E CG, Forzano F, Radojkovic D, Rial-Sebbag E, de Wert G, Borry P, Cornel MC; Public,and Professional Policy Committee of the European Society of Human Genetics. One small edit for humans, one giant edit for humankind? Points and questions to consider for a responsible way forward for gene editing in humans. Eur J Hum Genet, 2018,26(1):1-11. |
| [19] | Reeves RG, Voeneky S, Caetano-Anollés D, Beck F, Boëte C . Agricultural research, or a new bioweapon system? Science, 2018,362(6410):35-37. |
| [20] | Grunwald HA, Gantz VM, Poplawski G, Xu XS, Bier E, Cooper KL . Super-Mendelian inheritance mediated by CRISPR-Cas9 in the female mouse germline. Nature, 2019,566(7742):105-109. |
| [21] | Gangopadhyay SA, Cox KJ, Manna D, Lim D, Maji B, Zhou QX, Choudhary A . Precision control of CRISPR- Cas9 using small molecules and light. Biochemistry, 2019,58(4):234-244. |
| [22] | Maji B, Gangopadhyay SA, Lee M, Shi MC, Wu P, Heler R, Mok B, Lim D, Siriwardena SU, Paul B, Dančík V, Vetere A, Mesleh MF, Marraffini LA, Liu DR, Clemons PA, Wagner BK, Choudhary A . A high-throughput platform to identify small-molecule inhibitors of CRISPR-Cas9. Cell, 2019,177(4):1067-1079. |
| [23] | Hille F, Richter H, Wong SP, Bratovič M, Ressel S, Charpentier E . The biology of CRISPR-Cas: backward and forward. Cell, 2018,172(6):1239-1259. |
| [24] | Stanley SY, Maxwell KL . Phage-encoded anti-CRISPR defenses. Annu Rev Genet, 2018,52:445-464. |
| [25] | Zhang F, Song GX, Tian Y . Anti-CRISPRs: The natural inhibitors for CRISPR-Cas systems. Animal Model Exp Med, 2019,2(2):69-75. |
| [26] | Borges AL, Davidson AR, Bondy-Denomy J . The discovery, mechanisms, and evolutionary impact of anti-CRISPRs. Annu Rev Virol, 2017,4(1):37-59. |
| [27] | Pawluk A, Davidson AR, Maxwell KL . Anti-CRISPR: discovery, mechanism and function. Nat Rev Microbiol, 2018,16(1):12-17. |
| [28] | Davis KM, Pattanayak V, Thompson DB, Zuris JA, Liu DR . Small molecule-triggered Cas9 protein with improved genome-editing specificity. Nat Chem Biol, 2015,11(5):316-318. |
| [29] | Maji B, Moore CL, Zetsche B, Volz SE, Zhang F, Shoulders MD, Choudhary A . Multidimensional chemical control of CRISPR-Cas9. Nat Chem Biol, 2017,13(1):9-11. |
| [30] | Gao YC, Xiong X, Wong S, Charles EJ, Lim WA, Qi LS . Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat Methods, 2016,13(12):1043-1049. |
| [31] | Fegan A, White B, Carlson JC, Wagner CR . Chemically controlled protein assembly: techniques and applications. Chem Rev, 2010,110(6):3315-3336. |
| [32] | Chiarella AM, Butler KV, Gryder BE, Lu DB, Wang TA, Yu XF, Pomella S, Khan J, Jin J, Hathaway NA . Dose- dependent activation of gene expression is achieved using CRISPR and small molecules that recruit endogenous chromatin machinery. Nat Biotechnol, 2020,38(1):50-55. |
| [33] | Shrimp JH, Grose C, Widmeyer SRT, Thorpe AL, Jadhav A, Meier JL . Chemical control of a CRISPR-Cas9 acetyltransferase. ACS Chem Biol, 2018,13(2):455-460. |
| [34] | Rose JC, Stephany JJ, Valente WJ, Trevillian BM, Dang HV, Bielas JH, Maly DJ, Fowler DM . Rapidly inducible Cas9 and DSB-ddPCR to probe editing kinetics. Nat Methods, 2017,14(9):891-896. |
| [35] | Kang K, Huang L, Li Q, Liao XY, Dang QJ, Yang Y, Luo J, Zeng Y, Li L, Gou DM . An improved Tet-on system in microRNA overexpression and CRISPR/Cas9-mediated gene editing. J Anim Sci Biotechnol, 2019,10:43. |
| [36] | Nihongaki Y, Yamamoto S, Kawano F, Suzuki H, Sato M . CRISPR-Cas9-based photoactivatable transcription system. Chem Biol, 2015,22(2):169-174. |
| [37] | Nihongaki Y, Kawano F, Nakajima T, Sato M . Photoactivatable CRISPR-Cas9 for optogenetic genome editing. Nat Biotechnol, 2015,33(7):755-760. |
| [38] | Jain PK, Ramanan V, Schepers AG, Dalvie NS, Panda A, Fleming HE, Bhatia SN . Development of light-activated CRISPR using guide RNAs with photocleavable protectors. Angew Chem Int Ed Engl, 2016,55(40):12440-12444. |
| [39] | Tang WX, Hu JH, Liu DR . Aptazyme-embedded guide RNAs enable ligand-responsive genome editing and transcriptional activation. Nat Commun, 2017,8:15939. |
| [40] | Pu JY, Kentala K, Dickinson BC . Multidimensional control of Cas9 by evolved RNA Polymerase-Based biosensors. ACS Chem Biol, 2018,13(2):431-437. |
| [41] | Lai AC, Crews CM . Induced protein degradation: an emerging drug discovery paradigm. Nat Rev Drug Discov, 2017,16(2):101-114. |
| [42] | Natsume T, Kanemaki MT . Conditional degrons for controlling protein expression at the protein level. Annu Rev Genet, 2017,51:83-102. |
| [43] | Nabet B, Roberts JM, Buckley DL, Paulk J, Dastjerdi S, Yang A, Leggett AL, Erb MA, Lawlor MA, Souza A, Scott TG, Vittori S, Perry JA, Qi J, Winter GE, Wong KK, Gray NS, Bradner JE . The dTAG system for immediate and target-specific protein degradation. Nat Chem Biol, 2018,14(5):431-441. |
| [44] | Bonger KM, Chen LC, Liu CW, Wandless TJ . Small-molecule displacement of a cryptic degron causes conditional protein degradation. Nat Chem Biol, 2011,7(8):531-537. |
| [45] | Chung HK, Jacobs CL, Huo Y, Yang J, Krumm SA, Plemper RK, Tsien RY, Lin MZ . Tunable and reversible drug control of protein production via a self-excising degron. Nat Chem Biol, 2015,11(9):713-720. |
| [46] | Matsuzawa S, Cuddy M, Fukushima T, Reed JC . Method for targeting protein destruction by using a ubiquitin- independent, proteasome-mediated degradation pathway. Proc Natl Acad Sci USA, 2005,102(42):14982-14987. |
| [47] | Tu ZC, Yang WL, Yan S, Yin A, Gao JQ, Liu XD, Zheng YH, Zheng JZ, Li ZJ, Yang S, Li SH, Guo XY, Li XJ . Promoting Cas9 degradation reduces mosaic mutations in non-human primate embryos. Sci Rep, 2017,7:42081. |
| [48] | Charlesworth CT, Deshpande PS, Dever DP, Camarena J, Lemgart VT, Cromer MK, Vakulskas CA, Collingwood MA, Zhang LY, Bode NM, Behlke MA, Dejene B, Cieniewicz B, Romano R, Lesch BJ, Gomez-Ospina N, Mantri S, Pavel-Dinu M, Weinberg KI, Porteus MH . Identification of preexisting adaptive immunity to Cas9 proteins in humans. Nat Med, 2019,25(2):249-254. |
| [49] | Liu Q, Zhang HX, Huang XT . Anti-CRISPR proteins targeting the CRISPR-Cas system enrich the toolkit for genetic engineering. FEBS J, 2020,287(4):626-644. |
| [50] | Bondy-Denomy J, Davidson AR, Doudna JA, Fineran PC, Maxwell KL, Moineau S, Peng X, Sontheimer EJ, Wiedenheft B . A unified resource for tracking anti-CRISPR names. CRISPR J, 2018,1:304-305. |
| [51] | Zhu YL, Gao A, Zhan Q, Wang Y, Feng H, Liu SQ, Gao GX, Serganov A, Gao P . Diverse mechanisms of CRISPR- Cas9 inhibition by type IIC anti-CRISPR proteins. Mol Cell, 2019,74(2):296-309. |
| [52] | Jiang FG, Liu JJ, Osuna BA, Xu M, Berry JD, Rauch BJ, Nogales E, Bondy-Denomy J, Doudna JA . Temperature- responsive competitive inhibition of CRISPR-Cas9. Mol Cell, 2019,73(3):601-610. |
| [53] | Barkau CL, O'Reilly D, Rohilla KJ, Damha MJ, Gagnon KT,. Rationally designed anti-CRISPR nucleic acid inhibitors of CRISPR-Cas9. Nucleic Acid Ther, 2019,29(3):136-147. |
| [54] | Dowdy SF . Controlling CRISPR-Cas9 gene editing. N Engl J Med, 2019,381(3):289-290. |
| [55] | Lee J, Mou HW, Ibraheim R, Liang SQ, Liu PP, Xue W, Sontheimer EJ . Tissue-restricted genome editing in vivo specified by microRNA-repressible anti-CRISPR proteins. RNA, 2019,25(11):1421-1431. |
| [56] | Hoffmann MD, Aschenbrenner S, Grosse S, Rapti K, Domenger C, Fakhiri J, Mastel M, Börner K, Eils R, Grimm D, Niopek D . Cell-specific CRISPR-Cas9 activation by microRNA-dependent expression of anti-CRISPR proteins. Nucleic Acids Res, 2019,47(13):e75. |
| [57] | Wang SR, Wu LY, Huang HY, Xiong W, Liu J, Wei L, Yin P, Tian T, Zhou X . Conditional control of RNA-guided nucleic acid cleavage and gene editing. Nat Commun, 2020,11(1):91. |
| [1] | Zhong Bian, Dongping Cao, Wenshu Zhuang, Shuwei Zhang, Qiaoquan Liu, Lin Zhang. Revelation of rice molecular design breeding: the blend of tradition and modernity [J]. Hereditas(Beijing), 2023, 45(9): 718-740. |
| [2] | Bingzheng Wang, Chao Zhang, Jiali Zhang, Jin Sun. Conditional editing of the Drosophila melanogaster genome using single transcripts expressing Cas9 and sgRNA [J]. Hereditas(Beijing), 2023, 45(7): 593-601. |
| [3] | Xiaojun Zhang, Kun Xu, Juncen Shen, Lu Mu, Hongrun Qian, Jieyu Cui, Baoxia Ma, Zhilong Chen, Zhiying Zhang, Zehui Wei. A CRISPR/Cas9-Gal4BD donor adapting system for enhancing homology-directed repair [J]. Hereditas(Beijing), 2022, 44(8): 708-719. |
| [4] | Yuting Han, Bowen Xu, Yutong Li, Xinyi Lu, Xizhi Dong, Yuhao Qiu, Qinyun Che, Ruibao Zhu, Li Zheng, Xiaochen Li, Xu Si, Jianquan Ni. The cutting edge of gene regulation approaches in model organism Drosophila [J]. Hereditas(Beijing), 2022, 44(1): 3-14. |
| [5] | Chen Xuemei, Wei Yunlin, Ji Xiuling. Research progress of prophages [J]. Hereditas(Beijing), 2021, 43(3): 240-248. |
| [6] | Dingwei Peng, Ruiqiang Li, Wu Zeng, Min Wang, Xuan Shi, Jianhua Zeng, Xiaohong Liu, Yaoshen Chen, Zuyong He. Editing the cystine knot motif of MSTN enhances muscle development of Liang Guang Small Spotted pigs [J]. Hereditas(Beijing), 2021, 43(3): 261-270. |
| [7] | Guoling Li, Shanxin Yang, Zhenfang Wu, Xianwei Zhang. Recent developments in enhancing the efficiency of CRISPR/Cas9- mediated knock-in in animals [J]. Hereditas(Beijing), 2020, 42(7): 641-656. |
| [8] | Yingnan Chen, Jing Lu. Application of CRISPR/Cas9 mediated gene editing in trees [J]. Hereditas(Beijing), 2020, 42(7): 657-668. |
| [9] | Lianchao Tang, Feng Gu. Next-generation CRISPR-Cas for genome editing: focusing on the Cas protein and PAM [J]. Hereditas(Beijing), 2020, 42(3): 236-249. |
| [10] | Qianqian Zhang,Xuan Shang,Wanying Lin,Xiangmin Xu. Effect of genetic modifiers on the clinical severity of β-thalassemia [J]. Hereditas(Beijing), 2019, 41(8): 669-676. |
| [11] | Xuran Niu,Shuming Yin,Xi Chen,Tingting Shao,Dali Li. Gene editing technology and its recent progress in disease therapy [J]. Hereditas(Beijing), 2019, 41(7): 582-598. |
| [12] | Hao Ou,Guoling Li,Haoqiang Wang,Guangyan Huang,Gengyuan Cai,Zicong Li,Zhenfang Wu,Xianwei Zhang. MEK inhibitor PD0325901 significantly boosts ssODN-mediated HDR efficiency in porcine fetal fibroblasts [J]. Hereditas(Beijing), 2019, 41(4): 327-336. |
| [13] | Yunxiao Ren, Rudan Xiao, Xiaomin Lou, Xiangdong Fang. Research advance and application in the gene therapy of gene editing technologies [J]. Hereditas(Beijing), 2019, 41(1): 18-27. |
| [14] | Gaowei Xin, Xixun Hu, Kejian Wang, Xingchun Wang. Cas9 protein variant VQR recognizes NGAC protospacer adjacent motif in rice [J]. Hereditas(Beijing), 2018, 40(12): 1112-1119. |
| [15] | Yiou Chen, Ying Bao, Huazheng Ma, Zongyi Yi, Zhuo Zhou, Wensheng Wei. Gene editing technology and its research progress in China [J]. Hereditas(Beijing), 2018, 40(10): 900-915. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||