Hereditas(Beijing) ›› 2021, Vol. 43 ›› Issue (6): 615-622.doi: 10.16288/j.yczz.21-084
• Research Article • Previous Articles
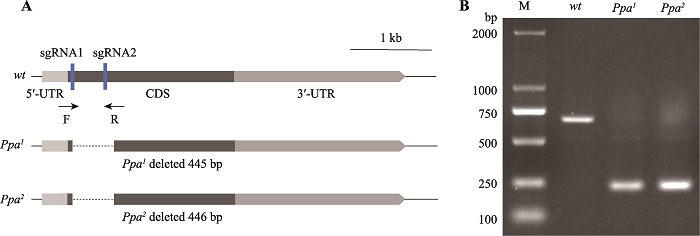
The F-box gene Ppa promotes lipid storage in Drosophila
- College of Life Sciences, Guizhou University, Guiyang 550025, China
-
Received:2021-03-05Revised:2021-04-28Online:2021-06-20Published:2021-05-07 -
Contact:Tian Yuan E-mail:ygw1996426@163.com;ytian1@gzu.edu.cn -
Supported by:Supported by the National Natural Science Foundation of China No(31600962);Guizhou Provincial Science and Technology Foundation No([2017]1043)
Cite this article
Guangwu Yang, Yuan Tian. The F-box gene Ppa promotes lipid storage in Drosophila[J]. Hereditas(Beijing), 2021, 43(6): 615-622.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
Table 1
The fly strains used in this study"
| 基因型 | 来源 | 用途 |
|---|---|---|
| w1118 | 中国科学院遗传与发育生物学研究所黄勋研究员实验室 | 对照组 |
| w*; ln(2LR)noc4LScorv9R, b1/CyO, P {ActGFP}JMR1 | 中国科学院遗传与发育生物学研究所黄勋研究员实验室 | 平衡致死系 |
| da-Gal4 | 清华大学果蝇中心 | 全身表达的Gal4 |
| y1M{nos-Cas9.P}ZH-2A w* | Bloomington | 表达Cas9蛋白 |
| w;L/CyO | 本实验室 | 平衡致死系 |
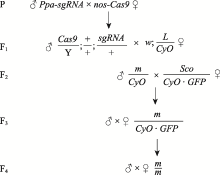
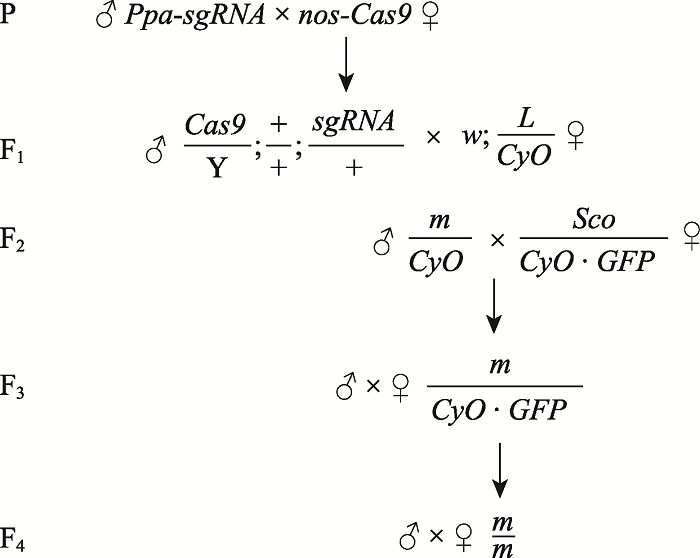
| Ppa-sgRNA | 本实验构建 | 表达两段sgRNA |
| UAS-Ppa-RA/RC | 本实验构建 | 超量表达PPA |
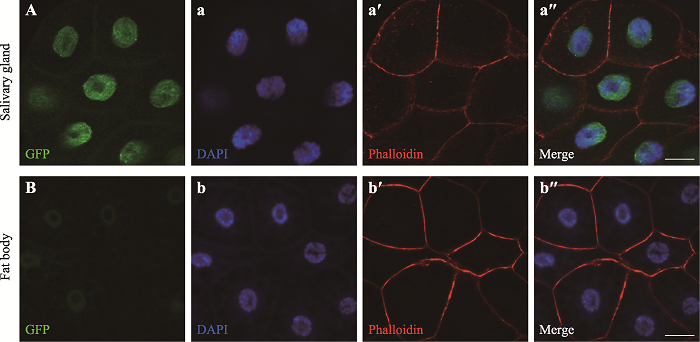
| UAS-Ppa-RA/RC-eGFP | 本实验构建 | PPA的亚细胞定位 |
Table 1
| [1] | Fahy E, Cotter D, Sud M, Subramaniam S. Lipid classification, structures and tools. Biochim Biophys Acta, 2011,1118(11):637-647. |
| [2] | Welte MA, Gould AP. Lipid droplet functions beyond energy storage. Biochim Biophys Acta Mol Cell Biol Lipids, 2017,1862(10 Pt B):1260-1272. |
| [3] | Welte MA. Expanding roles for lipid droplets. Curr Biol, 2015,25(11):R470-R481. |
| [4] | Filipe A, Mclauchlan J. Hepatitis C virus and lipid droplets: finding a niche. Trends Mol Med, 2015,21(1):34-42. |
| [5] | Inloes JM, Hsu KL, Dix MM, Viader A, Masuda K, Takei T, Wood MR, Cravatt BF. The hereditary spastic paraplegia-related enzyme DDHD2 is a principal brain triglyceride lipase. Proc Natl Acad Sci USA, 2014,111(41):14924-14929. |
| [6] | Xu X, Park JG, So JS, Lee AH. Transcriptional activation of Fsp27 by the liver-enriched transcription factor CREBH promotes lipid droplet growth and hepatic steatosis. Hepatology, 2015,61(3):857-869. |
| [7] | Wilfling F, Wang HJ, Haas JT, Krahmer N, Gould TJ, Uchida A, Cheng JX, Graham M, Christiano R, Fröhlich F, Liu XR, Buhman KK, Coleman RA, Bewersdorf J, Farese RV, Walther TC. Triacylglycerol synthesis enzymes mediate lipid droplet growth by relocalizing from the ER to lipid droplets. Dev Cell, 2013,24(4):384-399. |
| [8] | Krahmer N, Guo Y, Wilfling F, Hilger M, Lingrell S, Heger K, Newman HW, Schmidt-supprian M, Vance DE, Mann M, Farese RV, Walther TC. Phosphatidylcholine synthesis for lipid droplet expansion is mediated by localized activation of CTP:phosphocholine cytidylyltransferase. Cell Metab, 2011,14(4):504-515. |
| [9] | Wang XY, Sun LP, Zhang JF, Li H, Lv WQ, Zhang QQ. F-box proteins and their functions. Chin Bull Life Sci, 2008,20(5):807-811. |
| 王秀燕, 孙莉萍, 张建锋, 李辉, 吕文清, 张其清. F-box蛋白家族及其功能. 生命科学, 2008,20(5):807-811. | |
| [10] | Kipreos ET, Pagano M. The F-box protein family. Genome Biol, 2000, 1(5): REVIEWS3002. |
| [11] | Chen K, Cheng HH, Zhou RJ. Molecular mechanisms and functions of autophagy and the ubiq-uitin-proteasome pathway. Hereditas(Beijing), 2012,34(1):5-18. |
| 陈科, 程汉华, 周荣家. 自噬与泛素化蛋白降解途径的分子机制及其功能. 遗传, 2012,34(1):5-18. | |
| [12] | Xie CM, Wei WY, Sun Y. Role of SKP1-CUL1-F-box- protein (SCF) E3 ubiquitin ligases in skin cancer. J Genet Genomics, 2013,40(3):97-106. |
| [13] | Sohn JH, Lee YK, Han JS, Jeon YG, Kim JI, Choe SS, Kim SJ, Yoo HJ, Kim JB. Perilipin 1 (Plin1) deficiency promotes inflammatory responses in lean adipose tissue through lipid dysregulation. J Biol Chem, 2018,293(36):13974-13988. |
| [14] | Ju LP, Han JF, Zhang XY, Deng YJ, Yan H, Wang CR, Li XH, Chen SQ, Alimujiang M, Li X, Fang QC, Yang Y, Jia WP. Obesity-associated inflammation triggers an autophagy- lysosomal response in adipocytes and causes degradation of perilipin 1. Cell Death Dis, 2019,10(2):121. |
| [15] | Dui W, Lu W, Ma J, Jiao RJ. A systematic phenotypic screen of F-box genes through a tissue-specific RNAi- based approach in Drosophila. J Genet Genomics, 2012,39(8):397-413. |
| [16] | Clark E, Akam M. Odd-paired controls frequency doubling in Drosophila segmentation by altering the pair-rule gene regulatory network. eLife, 2016,5: e18215. |
| [17] | Raj L, Vivekanand P, Das TK, Badam E, Fernandes M, Finley RL, Brent R, Appel LF, Hanes SD, Weir M. Targeted localized degradation of Paired protein in Drosophila development. Curr Biol, 2000,10(20):1265-1272. |
| [18] | Heun P, Erhardt S, Blower MD, Weiss S, Skora AD, Karpen GH. Mislocalization of the Drosophila centromere- specific histone CID promotes formation of functional ectopic kinetochores. Dev Cell, 2006,10(3):303-315. |
| [19] | Moreno-moreno O, Medina-giró S, Torras-llort M, Azorín F. The F box protein partner of paired regulates stability of Drosophila centromeric histone H3, CenH3(CID). Curr Biol, 2011,21(17):1488-1493. |
| [20] | Zhao JJ, Xiong XL, Li Y, Liu X, Wang T, Zhang H, Jiao Y, Jiang JJ, Zhang HJ, Tang QQ, Gao X, Li XJ, Lu Y, Liu B, Hu C, Li XY. Hepatic F-box protein FBXW7 maintains glucose homeostasis through degradation of fetuin-A. Diabetes, 2018,67(5):818-830. |
| [21] | Zhang C, Chen F, Feng L, Shan Q, Zheng GH, Wang YJ, Lu J, Fan SH, Sun CH, Wu DM, Li MQ, Hu B, Wang QQ, Zhang ZF, Zheng YL. FBXW7 suppresses HMGB1- mediated innate immune signaling to attenuate hepatic inflammation and insulin resistance in a mouse model of nonalcoholic fatty liver disease. Mol Med, 2019,25(1):29. |
| [22] | Cooke PS, Holsberger DR, Cimafranca MA, Meling DD, Beals CM, Nakayama K, Nakayama KI, Kiyokawa H. The F box protein S phase kinase-associated protein 2 regulates adipose mass and adipocyte number in vivo. Obesity (Silver Spring), 2007,15(6):1400-1408. |
| [23] | Scott NA, Sharpe LJ, Capell-Hattam IM, Gullo SJ, Luu W, Brown AJ. The E3 ubiquitin ligase MARCHF6 as a metabolic integrator in cholesterol synthesis and beyond. Biochim Biophys Acta Mol Cell Biol Lipids, 2021,1866(1):158837. |
| [24] | Scott NA, Sharpe LJ, Capell-hattam IM, Gullo SJ, Luu W, Brown AJ. The cholesterol synthesis enzyme lanosterol 14α-demethylase is post-translationally regulated by the E3 ubiquitin ligase MARCH6. Biochem J, 2020,477(2):541-555. |
| [25] | Ghosh M, Niyogi S, Bhattacharyya M, Adak M, Nayak DK, Chakrabarti S, Chakrabarti P. Ubiquitin ligase COP1 controls hepatic fat metabolism by targeting ATGL for degradation. Diabetes, 2016,65(12):3561-3572. |
| [26] | Liu LL, Guo AW, Li QQ, Wu PF, Yang YJ, Chen FF, Li SH, Guo PJ, Zhang Q. The regulation of ubiquitination in milk fat synthesis in bovine. Hereditas(Beijing), 2020,42(6):548-555. |
| 刘莉莉, 郭爱伟, 李青青, 吴培福, 杨亚晋, 陈粉粉, 李素华, 郭盘江, 张勤. 泛素化途径在奶牛乳脂生成过程中的调控作用. 遗传, 2020,42(6):548-555. | |
| [27] | Leber V, Nans A, Singleton MR. Structural basis for assembly of the CBF3 kinetochore complex. EMBO J, 2018,37(2):269-281. |
| [1] | Bingzheng Wang, Chao Zhang, Jiali Zhang, Jin Sun. Conditional editing of the Drosophila melanogaster genome using single transcripts expressing Cas9 and sgRNA [J]. Hereditas(Beijing), 2023, 45(7): 593-601. |
| [2] | Meizhen Liu, Liren Wang, Yongmei Li, Xueyun Ma, Honghui Han, Dali Li. Generation of genetically modified rat models via the CRISPR/Cas9 technology [J]. Hereditas(Beijing), 2023, 45(1): 78-87. |
| [3] | Xiaojun Zhang, Kun Xu, Juncen Shen, Lu Mu, Hongrun Qian, Jieyu Cui, Baoxia Ma, Zhilong Chen, Zhiying Zhang, Zehui Wei. A CRISPR/Cas9-Gal4BD donor adapting system for enhancing homology-directed repair [J]. Hereditas(Beijing), 2022, 44(8): 708-719. |
| [4] | Chong Zhang, Zixuan Wei, Min Wang, Yaosheng Chen, Zuyong He. Editing MC1R in human melanoma cells by CRISPR/Cas9 and functional analysis [J]. Hereditas(Beijing), 2022, 44(7): 581-590. |
| [5] | Yao Liu, Xianhui Zhou, Shuhong Huang, Xiaolong Wang. Prime editing: a search and replace tool with versatile base changes [J]. Hereditas(Beijing), 2022, 44(11): 993-1008. |
| [6] | Yuting Han, Bowen Xu, Yutong Li, Xinyi Lu, Xizhi Dong, Yuhao Qiu, Qinyun Che, Ruibao Zhu, Li Zheng, Xiaochen Li, Xu Si, Jianquan Ni. The cutting edge of gene regulation approaches in model organism Drosophila [J]. Hereditas(Beijing), 2022, 44(1): 3-14. |
| [7] | Dingwei Peng, Ruiqiang Li, Wu Zeng, Min Wang, Xuan Shi, Jianhua Zeng, Xiaohong Liu, Yaoshen Chen, Zuyong He. Editing the cystine knot motif of MSTN enhances muscle development of Liang Guang Small Spotted pigs [J]. Hereditas(Beijing), 2021, 43(3): 261-270. |
| [8] | Na Wang, Zhilian Jia, Qiang Wu. RFX5 regulates gene expression of the Pcdhα cluster [J]. Hereditas(Beijing), 2020, 42(8): 760-774. |
| [9] | Guoling Li, Shanxin Yang, Zhenfang Wu, Xianwei Zhang. Recent developments in enhancing the efficiency of CRISPR/Cas9- mediated knock-in in animals [J]. Hereditas(Beijing), 2020, 42(7): 641-656. |
| [10] | Yingnan Chen, Jing Lu. Application of CRISPR/Cas9 mediated gene editing in trees [J]. Hereditas(Beijing), 2020, 42(7): 657-668. |
| [11] | Siyuan Liu, Guoqiang Yi, Zhonglin Tang, Bin Chen. Progress on genome-wide CRISPR/Cas9 screening for functional genes and regulatory elements [J]. Hereditas(Beijing), 2020, 42(5): 435-443. |
| [12] | Liwen Bao, Yiye Zhou, Fanyi Zeng. Advances in gene therapy for β-thalassemia and hemophilia based on the CRISPR/Cas9 technology [J]. Hereditas(Beijing), 2020, 42(10): 949-964. |
| [13] | Minting Lin, Lulu Lai, Miao Zhao, Biwei Lin, Xiangping Yao. Construction of a striatum-specific Slc20a2 gene knockout mice model by CRISPR/Cas9 AAV system [J]. Hereditas(Beijing), 2020, 42(10): 1017-1027. |
| [14] | Peifeng Liu, Qiang Wu. Probing 3D genome by CRISPR/Cas9 [J]. Hereditas(Beijing), 2020, 42(1): 18-31. |
| [15] | Jue Wang, Juan Huang, Rui Xu. Seamless genome editing in Drosophila by combining CRISPR/Cas9 and piggyBac technologies [J]. Hereditas(Beijing), 2019, 41(5): 422-429. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||