Hereditas(Beijing) ›› 2020, Vol. 42 ›› Issue (8): 760-774.doi: 10.16288/j.yczz.20-184
• Research Article • Previous Articles Next Articles
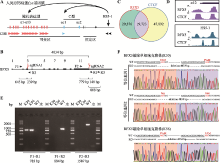
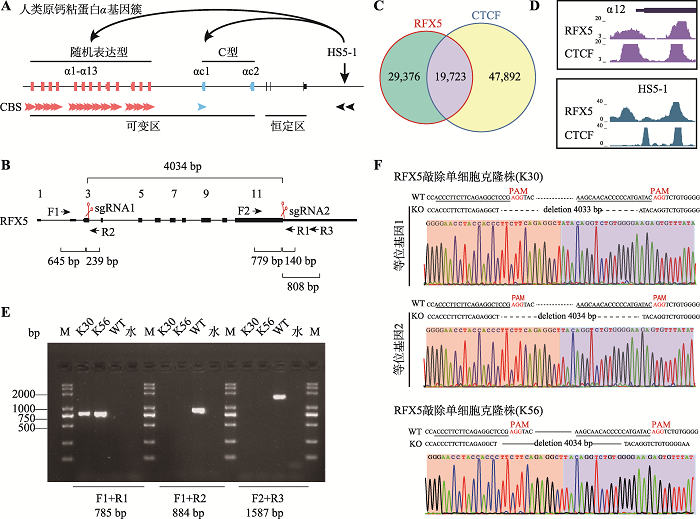
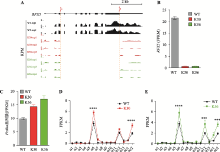
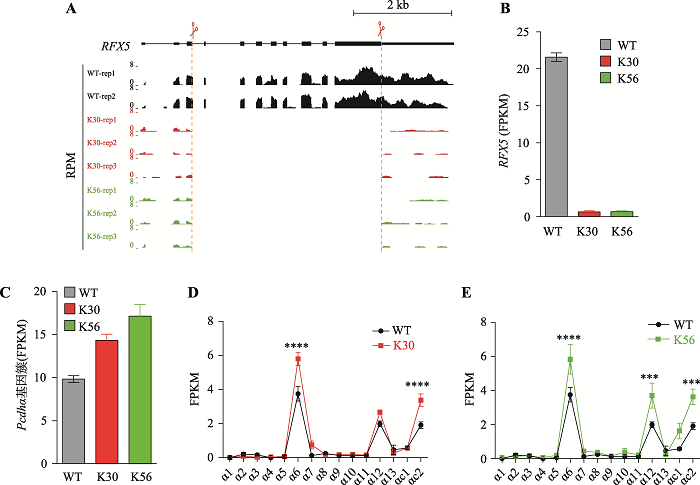
RFX5 regulates gene expression of the Pcdhα cluster
Na Wang, Zhilian Jia, Qiang Wu( )
)
- Center for Comparative Biomedicine, Key laboratory of Systems Biomedicine (Ministry of Education), Institute of Systems Biomedicine, Shanghai Jiao Tong University, Shanghai 200240, China
-
Received:2020-06-19Revised:2020-07-08Online:2020-08-20Published:2020-07-29 -
Contact:Wu Qiang E-mail:qwu123@gmail.com -
Supported by:Supported by the National Natural Science Foundation of China Nos(91640118);Supported by the National Natural Science Foundation of China Nos(31630039)
Cite this article
Na Wang, Zhilian Jia, Qiang Wu. RFX5 regulates gene expression of the Pcdhα cluster[J]. Hereditas(Beijing), 2020, 42(8): 760-774.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
Table 1
Primer sequences used in this study"
| 类型 | 引物名称 | 序列(5′→3′) |
|---|---|---|
| PCR | F1 | GGGAAGTCGTGGCGAGATTA |
| F2 | GTGCCCTGAAAGTGGCTACA | |
| R1 | CTGGGTGACTCAGCTGTCTG | |
| R2 | GCTGGAGGTCACACACAAGA | |
| R3 | AGAGTAGCCTGGTGTTAGCG | |
| sgRNA | sgRNA1F | ACCGACCCTTCTTCAGAGGCTCCG |
| sgRNA1R | AAACCGGAGCCTCTGAAGAAGGGT | |
| sgRNA2F | ACCGAAGCAACACCCCCATGATAC | |
| sgRNA2R | AAACGTATCATGGGGGTGTTGCTT | |
| Index | P5-HS5-1-F2 | AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCTGTTTTGGCGGCGACAAATTCG |
| P5-α12-F3 | AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCTAGTCCAATCATTCACGGAATAGGATC | |
| P7-index-1 | CAAGCAGAAGACGGCATACGAGATCGAGTAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-2 | CAAGCAGAAGACGGCATACGAGATTCTCCGGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-3 | CAAGCAGAAGACGGCATACGAGATAATGAGGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-4 | CAAGCAGAAGACGGCATACGAGATGGAATCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-5 | CAAGCAGAAGACGGCATACGAGATAGCTTCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-6 | CAAGCAGAAGACGGCATACGAGATGCGCATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-7 | CAAGCAGAAGACGGCATACGAGATGCGCGAGAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-8 | CAAGCAGAAGACGGCATACGAGATAGAGTACTGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-9 | CAAGCAGAAGACGGCATACGAGATGCTCCGTAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-10 | CAAGCAGAAGACGGCATACGAGATCATGAGAGGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-11 | CAAGCAGAAGACGGCATACGAGATTGAATCGCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-12 | CAAGCAGAAGACGGCATACGAGATGTCGCGTAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-13 | CAAGCAGAAGACGGCATACGAGATCCGCATGAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-14 | CAAGCAGAAGACGGCATACGAGATCCACAATCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-15 | CAAGCAGAAGACGGCATACGAGATTTCATCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-16 | CAAGCAGAAGACGGCATACGAGATTAGAGTGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-17 | CAAGCAGAAGACGGCATACGAGATTAACTCGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-18 | CAAGCAGAAGACGGCATACGAGATTATACGGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-19 | CAAGCAGAAGACGGCATACGAGATGAGTCAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-20 | CAAGCAGAAGACGGCATACGAGATGACATAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-21 | CAAGCAGAAGACGGCATACGAGATGTATTAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-22 | CAAGCAGAAGACGGCATACGAGATGACACAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-23 | CAAGCAGAAGACGGCATACGAGATGCGATAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-24 | CAAGCAGAAGACGGCATACGAGATCTCTATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT | |
| P7-index-25 | CAAGCAGAAGACGGCATACGAGATTACGCAGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT |
| [1] |
Bickmore WA . The spatial organization of the human genome. Annu Rev Genomics Hum Genet, 2013,14:67-84.
doi: 10.1146/annurev-genom-091212-153515 pmid: 23875797 |
| [2] |
Wu Q, Maniatis T . A striking organization of a large family of human neural cadherin-like cell adhesion genes. Cell, 1999,97(6):779-790.
doi: 10.1016/s0092-8674(00)80789-8 pmid: 10380929 |
| [3] |
Wu Q . Comparative genomics and diversifying selection of the clustered vertebrate protocadherin genes. Genetics, 2005,169(4):2179-2188.
doi: 10.1534/genetics.104.037606 pmid: 15744052 |
| [4] |
Suo L, Lu HN, Ying GX, Capecchi MR, Wu Q . Protocadherin clusters and cell adhesion kinase regulate dendrite complexity through rho GTPase. J Mol Cell Biol, 2012,4(6):362-376.
doi: 10.1093/jmcb/mjs034 |
| [5] |
Wu Q, Maniatis T . Large exons encoding multiple ectodomains are a characteristic feature of protocadherin genes. Proc Natl Acad Sci USA, 2000,97(7):3124-3129.
doi: 10.1073/pnas.060027397 pmid: 10716726 |
| [6] |
Wu Q, Zhang T, Cheng JF, Kim Y, Grimwood J, Schmutz J, Dickson M, Noonan JP, Zhang MQ, Myers RM, Maniatis T . Comparative DNA sequence analysis of mouse and human protocadherin gene clusters. Genome Res, 2001,11(3):389-404.
doi: 10.1101/gr.167301 pmid: 11230163 |
| [7] |
Zou C, Huang W, Ying G, Wu Q . Sequence analysis and expression mapping of the rat clustered protocadherin gene repertoires. Neuroscience, 2007,144(2):579-603.
doi: 10.1016/j.neuroscience.2006.10.011 pmid: 17110050 |
| [8] |
Zhang T, Haws P, Wu Q . Multiple variable first exons: A mechanism for cell- and tissue-specific gene regulation. Genome Res, 2004,14(1):79-89.
doi: 10.1101/gr.1225204 pmid: 14672974 |
| [9] | Zhai YN, Xu Q, Guo Y, Wu Q . Characterization of a cluster of CTCF-binding sites in a protocadherin regulatory region. Hereditas(Beijing), 2016,38(4):323-336. |
| 翟亚男, 许泉, 郭亚, 吴强 . 原钙粘蛋白基因簇调控区域中成簇的CTCF结合位点分析. 遗传, 2016,38(4):323-336. | |
| [10] |
Tasic B, Nabholz CE, Baldwin KK, Kim Y, Rueckert EH, Ribich SA, Cramer P, Wu Q, Axel R, Maniatis T . Promoter choice determines splice site selection in protocadherin α and γ pre-mRNA splicing. Mol Cell, 2002,10(1):21-33.
doi: 10.1016/s1097-2765(02)00578-6 pmid: 12150904 |
| [11] |
Chen WV, Maniatis T . Clustered protocadherins. Development, 2013,140(16):3297-3302.
doi: 10.1242/dev.090621 |
| [12] |
Ribich S, Tasic B, Maniatis T . Identification of long-range regulatory elements in the protocadherin- α gene cluster. Proc Natl Acad Sci USA, 2006,103(52):19719-19724.
doi: 10.1073/pnas.0609445104 pmid: 17172445 |
| [13] |
Guo Y, Monahan K, Wu HY, Gertz J, Varley KE, Li W, Myers RM, Maniatis T, Wu Q . CTCF/cohesin-mediated DNA looping is required for protocadherin α promoter choice. Proc Natl Acad Sci USA, 2012,109(51):21081-21086.
pmid: 23204437 |
| [14] |
Guo Y, Xu Q, Canzio D, Shou J, Li JH, Gorkin DU, Jung I, Wu HY, Zhai YA, Tang YX, Lu YC, Wu YH, Jia ZL, Li W, Zhang MQ, Ren B, Krainer AR, Maniatis T, Wu Q . CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell, 2015,162(4):900-910.
doi: 10.1016/j.cell.2015.07.038 pmid: 26276636 |
| [15] |
Jia ZL, Li JW, Ge X, Wu YH, Guo Y, Wu Q . Tandem CTCF sites function as insulators to balance spatial chromatin contacts and topological enhancer-promoter selection. Genome Biol, 2020,21(1):75.
doi: 10.1186/s13059-020-01984-7 pmid: 32293525 |
| [16] |
Kagey MH, Newman JJ, Bilodeau S, Zhan Y, Orlando DA, van Berkum NL, Ebmeier CC, Goossens J, Rahl PB, Levine SS, Taatjes DJ, Dekker J, Young RA. Mediator and cohesin connect gene expression and chromatin architecture. Nature, 2010,467(7314):430-435.
doi: 10.1038/nature09380 pmid: 20720539 |
| [17] |
Emery P, Durand B, Mach B, Reith W . RFX proteins, a novel family of DNA binding proteins conserved in the eukaryotic kingdom. Nucleic Acids Res, 1996,24(5):803-807.
doi: 10.1093/nar/24.5.803 pmid: 8600444 |
| [18] |
Gajiwala KS, Chen H, Cornille F, Roques BP, Reith W, Mach B, Burley SK . Structure of the winged-helix protein hRFX1 reveals a new mode of DNA binding. Nature, 2000,403(6772):916-921.
doi: 10.1038/35002634 pmid: 10706293 |
| [19] |
Reith W, Mach B . The bare lymphocyte syndrome and the regulation of MHC expression. Annu Rev Immunol, 2001,19:331-373.
doi: 10.1146/annurev.immunol.19.1.331 pmid: 11244040 |
| [20] |
Mach B, Steimle V, Martinez-Soria E, Reith W . Regulation of MHC class II genes: Lessons from a disease. Annu Rev Immunol, 1996,14:301-331.
doi: 10.1146/annurev.immunol.14.1.301 pmid: 8717517 |
| [21] |
Klein C, Lisowska-Grospierre B, LeDeist F, Fischer A, Griscelli C. Major histocompatibility complex class II deficiency: Clinical manifestations, immunologic features, and outcome. J Pediatr, 1993,123(6):921-928.
doi: 10.1016/s0022-3476(05)80388-9 pmid: 8229525 |
| [22] |
Villard J, Masternak K, Lisowska-Grospierre B, Fischer A, Reith W . MHC class II deficiency: A disease of gene regulation. Medicine (Baltimore), 2001,80(6):405-418.
doi: 10.1097/00005792-200111000-00006 |
| [23] |
Griscelli C, Lisowska-Grospierre B, Mach B . Combined immunodeficiency with defective expression in MHC class II genes. Immunodefic Rev, 1989,1(2):135-153.
pmid: 2517209 |
| [24] |
Steimle V, Durand B, Barras EZ, Zufferey M, Hadam MR, Mach B, Reith W . A novel DNA-binding regulatory factor is mutated in primary MHC class II deficiency (bare lymphocyte syndrome). Genes Dev, 1995,9(9):1021-1032.
doi: 10.1101/gad.9.9.1021 pmid: 7744245 |
| [25] |
Guardiola J, Maffei A . Control of MHC class II gene expression in autoimmune, infectious, and neoplastic diseases. Crit Rev Immunol, 1993,13(3-4):247-268.
pmid: 8110378 |
| [26] |
Hanna S, Etzioni A . MHC class I and II deficiencies. J Allergy Clin Immunol, 2014,134(2):269-275.
doi: 10.1016/j.jaci.2014.06.001 pmid: 25001848 |
| [27] |
Cresswell P . Assembly, transport, and function of MHC class II molecules. Annu Rev Immunol, 1994,12:259-293.
doi: 10.1146/annurev.iy.12.040194.001355 pmid: 8011283 |
| [28] |
Villard J, Reith W, Barras E, Gos A, Morris MA, Antonarakis SE, Van den Elsen PJ, Mach B,. Analysis of mutations and chromosomal localisation of the gene encoding RFX5, a novel transcription factor affected in major histocompatibility complex class II deficiency. Hum Mutat, 1997,10(6):430-435.
doi: 10.1002/(SICI)1098-1004(1997)10:6<430::AID-HUMU3>3.0.CO;2-H pmid: 9401005 |
| [29] |
Boss JM . A common set of factors control the expression of the MHC class II, invariant chain, and HLA-DM genes. Microbes Infect, 1999,1(11):847-853.
doi: 10.1016/s1286-4579(99)00234-8 pmid: 10614001 |
| [30] |
Clausen BE, Waldburger JM, Schwenk F, Barras E, Mach B, Rajewsky K, Forster I, Reith W . Residual MHC class II expression on mature dendritic cells and activated B cells in RFX5-deficient mice. Immunity, 1998,8(2):143-155.
doi: 10.1016/s1074-7613(00)80467-7 pmid: 9491996 |
| [31] |
Villard J, Peretti M, Masternak K, Barras E, Caretti G, Mantovani R, Reith W . A functionally essential domain of RFX5 mediates activation of major histocompatibility complex class II promoters by promoting cooperative binding between RFX and NF-Y. Mol Cell Biol, 2000,20(10):3364-3376.
doi: 10.1128/mcb.20.10.3364-3376.2000 pmid: 10779326 |
| [32] |
Stavride P, Arampatzi P, Papamatheakis J . Differential regulation of MHC II genes by PRMT6, via an AT-hook motif of RFX5. Mol Immunol, 2013,56(4):390-398.
doi: 10.1016/j.molimm.2013.05.235 |
| [33] |
Garvie CW, Stagno JR, Reid S, Singh A, Harrington E, Boss JM . Characterization of the RFX complex and the RFX5(L66A) mutant: Implications for the regulation of MHC class II gene expression. Biochemistry, 2007,46(6):1597-1611.
doi: 10.1021/bi6023868 pmid: 17279624 |
| [34] |
DeSandro AM, Nagarajan UM, Boss JM . Associations and interactions between bare lymphocyte syndrome factors. Mol Cell Biol, 2000,20(17):6587-6599.
doi: 10.1128/mcb.20.17.6587-6599.2000 pmid: 10938133 |
| [35] |
Laird KM, Briggs LL, Boss JM, Summers MF, Garvie CW . Solution structure of the heterotrimeric complex between the interaction domains of RFX5 and RFXAP from the RFX gene regulatory complex. J Mol Biol, 2010,403(1):40-51.
doi: 10.1016/j.jmb.2010.08.025 pmid: 20732328 |
| [36] |
Nagarajan UM, Long AB, Harreman MT, Corbett AH, Boss JM . A hierarchy of nuclear localization signals governs the import of the regulatory factor X complex subunits and MHC class II expression. J Immunol, 2004,173(1):410-419.
doi: 10.4049/jimmunol.173.1.410 pmid: 15210800 |
| [37] |
Sengupta PK, Fargo J, Smith BD . The RFX family interacts at the collagen (COL1A2) start site and represses transcription. J Biol Chem, 2002,277(28):24926-24937.
doi: 10.1074/jbc.M111712200 pmid: 11986307 |
| [38] |
Michel S, Linnebacher M, Alcaniz J, Voss M, Wagner R, Dippold W, Becker C, Doeberitz MV, Ferrone S, Kloor M . Lack of HLA class II antigen expression in microsatellite unstable colorectal carcinomas is caused by mutations in HLA class II regulatory genes. Int J Cancer, 2010,127(4):889-898.
doi: 10.1002/ijc.25106 pmid: 20013806 |
| [39] |
Zhao YJ, Xie XW, Liao WJ, Zhang HH, Cao H, Fei R, Wang XY, Wei L, Shao QX, Chen HS . The transcription factor RFX5 is a transcriptional activator of the TPP1 gene in hepatocellular carcinoma. Oncol Rep, 2017,37(1):289-296.
doi: 10.3892/or.2016.5240 pmid: 27840983 |
| [40] |
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E . A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science, 2012,337(6096):816-821.
doi: 10.1126/science.1225829 pmid: 22745249 |
| [41] |
Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, Boyaval P, Fremaux C, Horvath P, Magadan AH, Moineau S . The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature, 2010,468(7320):67-71.
doi: 10.1038/nature09523 pmid: 21048762 |
| [42] |
Li JH, Shou J, Guo Y, Tang YX, Wu YH, Jia ZL, Zhai YN, Chen ZF, Xu Q, Wu Q . Efficient inversions and duplications of mammalian regulatory DNA elements and gene clusters by CRISPR/Cas9. J Mol Cell Biol, 2015,7(4):284-298.
doi: 10.1093/jmcb/mjv016 pmid: 25757625 |
| [43] |
Shou J, Li JH, Liu YB, Wu Q . Precise and predictable CRISPR chromosomal rearrangements reveal principles of Cas9-mediated nucleotide insertion. Mol Cell, 2018,71(4):498-509.
doi: 10.1016/j.molcel.2018.06.021 pmid: 30033371 |
| [44] |
Shi X, Shou J, Mehryar MM, Li JW, Wang LY, Zhang M, Huang HY, Sun XF, Wu Q . Cas9 has no exonuclease activity resulting in staggered cleavage with overhangs and predictable di- and tri-nucleotide CRISPR insertions without template donor. Cell Discov, 2019,5:53.
doi: 10.1038/s41421-019-0120-z pmid: 31636963 |
| [45] |
Cong L, Ran FA, Cox D, Lin SL, Barretto R, Habib N, Hsu PD, Wu XB, Jiang WY, Marraffini LA, Zhang F . Multiplex genome engineering using CRISPR/Cas systems. Science, 2013,339(6121):819-823.
doi: 10.1126/science.1229223 |
| [46] | Li JH, Shou J, Wu Q . DNA fragment editing of genomes by CRISPR/Cas9. Hereditas(Beijing), 2015,37(10):992-1002. |
| 李金环, 寿佳, 吴强 . CRISPR/Cas9系统在基因组DNA片段编辑中的应用. 遗传, 2015,37(10):992-1002. | |
| [47] | Liu PF, Wu Q . Probing 3D genome by CRISPR/Cas9. Hereditas(Beijing), 2020,42(1):18-31. |
| 刘沛峰, 吴强 . CRISPR/Cas9基因编辑在三维基因组研究中的应用. 遗传, 2020,42(1):18-31. | |
| [48] |
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L . Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc, 2012,7(3):562-578.
doi: 10.1038/nprot.2012.016 |
| [49] |
He QY, Johnston J, Zeitlinger J . ChIP-nexus enables improved detection of in vivo transcription factor binding footprints. Nat Biotechnol, 2015,33(4):395-401.
doi: 10.1038/nbt.3121 pmid: 25751057 |
| [50] |
Hu JZ, Meyers RM, Dong JC, Panchakshari RA, Alt FW, Frock RL . Detecting DNA double-stranded breaks in mammalian genomes by linear amplification-mediated high-throughput genome-wide translocation sequencing. Nat Protoc, 2016,11(5):853-871.
doi: 10.1038/nprot.2016.043 pmid: 27031497 |
| [51] |
Filippova GN, Fagerlie S, Klenova EM, Myers C, Dehner Y, Goodwin G, Neiman PE, Collins SJ, Lobanenkov VV . An exceptionally conserved transcriptional repressor, CTCF, employs different combinations of zinc fingers to bind diverged promoter sequences of avian and mammalian c-myc oncogenes. Mol Cell Biol, 1996,16(6):2802-2813.
doi: 10.1128/mcb.16.6.2802 pmid: 8649389 |
| [52] |
Hadjur S, Williams LM, Ryan NK, Cobb BS, Sexton T, Fraser P, Fisher AG, Merkenschlager M . Cohesins form chromosomal cis-interactions at the developmentally regulated IFNG locus. Nature, 2009,460(7253):410-413.
doi: 10.1038/nature08079 pmid: 19458616 |
| [53] |
Wendt KS, Yoshida K, Itoh T, Bando M, Koch B, Schirghuber E, Tsutsumi S, Nagae G, Ishihara K, Mishiro T, Yahata K, Imamoto F, Aburatani H, Nakao M, Imamoto N, Maeshima K, Shirahige K, Peters JM . Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature, 2008,451(7180):796-801.
doi: 10.1038/nature06634 pmid: 18235444 |
| [54] |
Shi ZB, Gao HS, Bai XC, Yu HT . Cryo-EM structure of the human cohesin-NIPBL-DNA complex. Science, 2020,368(6498):1454-1459.
doi: 10.1126/science.abb0981 pmid: 32409525 |
| [55] |
Pugacheva EM, Kubo N, Loukinov D, Tajmul M, Kang S, Kovalchuk AL, Strunnikov AV, Zentner GE, Ren B, Lobanenkov VV . CTCF mediates chromatin looping via N-terminal domain-dependent cohesin retention. Proc Natl Acad Sci USA, 2020,117(4):2020-2031.
doi: 10.1073/pnas.1911708117 pmid: 31937660 |
| [56] | Wu HY, Guo Y, Li W, Wu Q . Cloning and functional analysis of the regulatory elements in the human protocadherin gene cluster. Life Sci Res, 2014,18(2):95-99. |
| 吴海洋, 郭亚, 李伟, 吴强 . 人类原钙粘蛋白基因簇调控元件的克隆及对其启动子活性的影响. 生命科学研究, 2014,18(2):95-99. | |
| [57] |
Kehayova P, Monahan K, Chen WS, Maniatis T . Regulatory elements required for the activation and repression of the protocadherin- α gene cluster. Proc Natl Acad Sci USA, 2011,108(41):17195-17200.
doi: 10.1073/pnas.1114357108 pmid: 21949399 |
| [58] |
Monahan K, Rudnick ND, Kehayova PD, Pauli F, Newberry KM, Myers RM, Maniatis T . Role of CCCTC binding factor (CTCF) and cohesin in the generation of single-cell diversity of protocadherin-α gene expression. Proc Natl Acad Sci USA, 2012,109(23):9125-9130.
doi: 10.1073/pnas.1205074109 pmid: 22550178 |
| [1] | Bingzheng Wang, Chao Zhang, Jiali Zhang, Jin Sun. Conditional editing of the Drosophila melanogaster genome using single transcripts expressing Cas9 and sgRNA [J]. Hereditas(Beijing), 2023, 45(7): 593-601. |
| [2] | Meizhen Liu, Liren Wang, Yongmei Li, Xueyun Ma, Honghui Han, Dali Li. Generation of genetically modified rat models via the CRISPR/Cas9 technology [J]. Hereditas(Beijing), 2023, 45(1): 78-87. |
| [3] | Xiaojun Zhang, Kun Xu, Juncen Shen, Lu Mu, Hongrun Qian, Jieyu Cui, Baoxia Ma, Zhilong Chen, Zhiying Zhang, Zehui Wei. A CRISPR/Cas9-Gal4BD donor adapting system for enhancing homology-directed repair [J]. Hereditas(Beijing), 2022, 44(8): 708-719. |
| [4] | Chong Zhang, Zixuan Wei, Min Wang, Yaosheng Chen, Zuyong He. Editing MC1R in human melanoma cells by CRISPR/Cas9 and functional analysis [J]. Hereditas(Beijing), 2022, 44(7): 581-590. |
| [5] | Yao Liu, Xianhui Zhou, Shuhong Huang, Xiaolong Wang. Prime editing: a search and replace tool with versatile base changes [J]. Hereditas(Beijing), 2022, 44(11): 993-1008. |
| [6] | Yuting Han, Bowen Xu, Yutong Li, Xinyi Lu, Xizhi Dong, Yuhao Qiu, Qinyun Che, Ruibao Zhu, Li Zheng, Xiaochen Li, Xu Si, Jianquan Ni. The cutting edge of gene regulation approaches in model organism Drosophila [J]. Hereditas(Beijing), 2022, 44(1): 3-14. |
| [7] | Cong Zhou, Qiangwei Zhou, Sheng Cheng, Guoliang Li. Research progress of CTCF in mediating 3D genome formation and regulating gene expression [J]. Hereditas(Beijing), 2021, 43(9): 816-821. |
| [8] | Xianglong He, Jinhuan Li, Qiang Wu. Combinatorial CRISPR inversions of CTCF sites in HOXD cluster reveal complex insulator function [J]. Hereditas(Beijing), 2021, 43(8): 758-774. |
| [9] | Ling Wang, Jinhuan Li, Haiyan Huang, Qiang Wu. Serial deletions of tandem reverse CTCF sites reveal balanced HOXD regulatory landscape of enhancers [J]. Hereditas(Beijing), 2021, 43(8): 775-791. |
| [10] | Guangwu Yang, Yuan Tian. The F-box gene Ppa promotes lipid storage in Drosophila [J]. Hereditas(Beijing), 2021, 43(6): 615-622. |
| [11] | Ye Wei, Ke Li, Daru Lu, Huaxing Zhu. Optimization of CUT&Tag product recovery and library construction method [J]. Hereditas(Beijing), 2021, 43(4): 362-374. |
| [12] | Dingwei Peng, Ruiqiang Li, Wu Zeng, Min Wang, Xuan Shi, Jianhua Zeng, Xiaohong Liu, Yaoshen Chen, Zuyong He. Editing the cystine knot motif of MSTN enhances muscle development of Liang Guang Small Spotted pigs [J]. Hereditas(Beijing), 2021, 43(3): 261-270. |
| [13] | Guoling Li, Shanxin Yang, Zhenfang Wu, Xianwei Zhang. Recent developments in enhancing the efficiency of CRISPR/Cas9- mediated knock-in in animals [J]. Hereditas(Beijing), 2020, 42(7): 641-656. |
| [14] | Yingnan Chen, Jing Lu. Application of CRISPR/Cas9 mediated gene editing in trees [J]. Hereditas(Beijing), 2020, 42(7): 657-668. |
| [15] | Siyuan Liu, Guoqiang Yi, Zhonglin Tang, Bin Chen. Progress on genome-wide CRISPR/Cas9 screening for functional genes and regulatory elements [J]. Hereditas(Beijing), 2020, 42(5): 435-443. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||