Hereditas(Beijing) ›› 2023, Vol. 45 ›› Issue (6): 488-500.doi: 10.16288/j.yczz.23-044
• Review • Previous Articles Next Articles
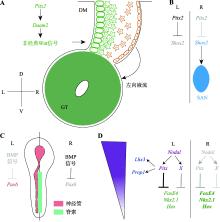
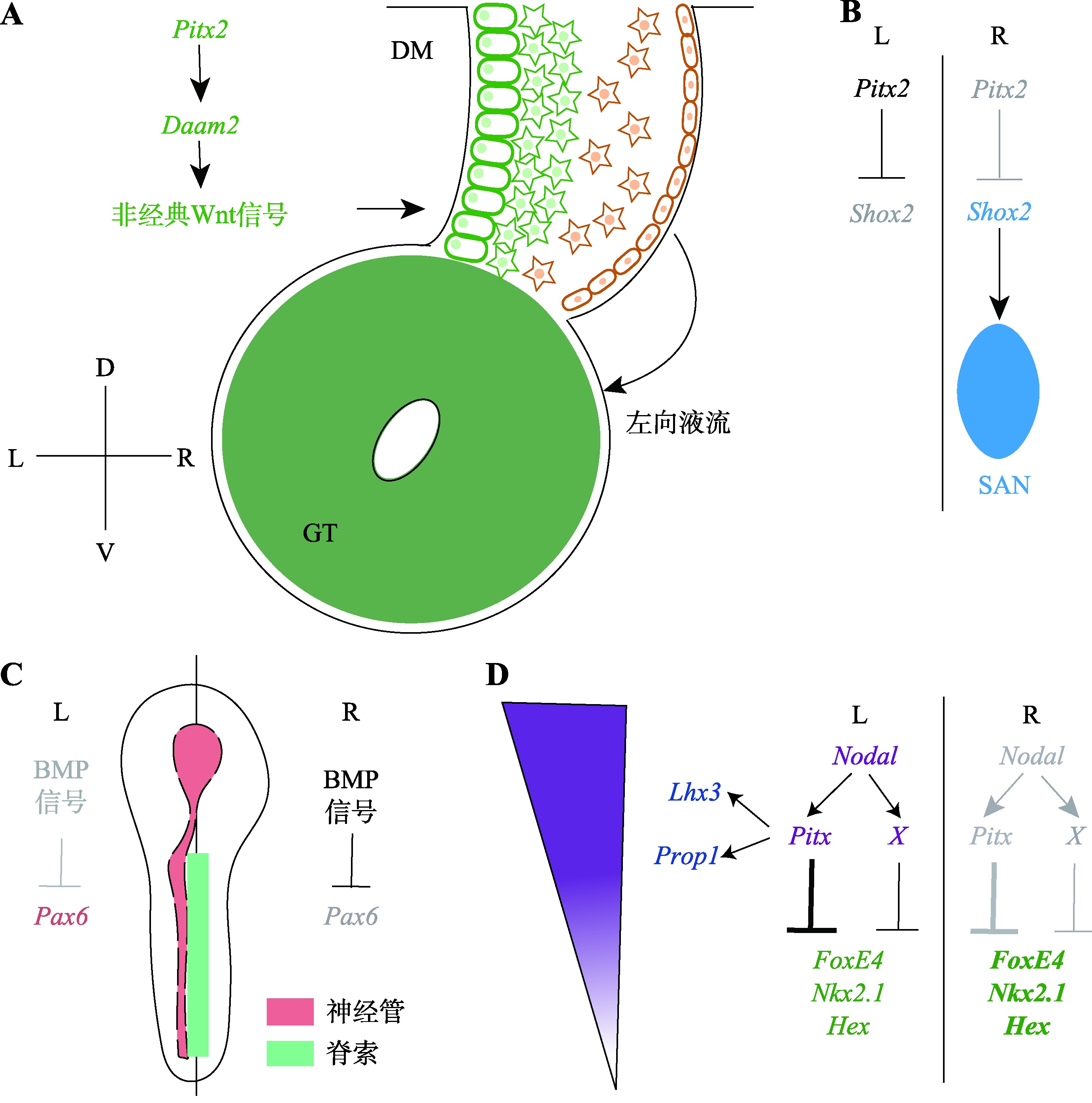
Progress on the mechanism of left-right asymmetrical patterning in bilaterians
Chaofan Xing1,2( ), Mintao Wang1,2, Lei Wang1,2, Xin Shen1,2
), Mintao Wang1,2, Lei Wang1,2, Xin Shen1,2
- 1. Jiangsu Key Laboratory of Marine Bioresources and Environment /Jiangsu Key Laboratory of Marine Biotechnology School of Marine Science and Fisheries, Jiangsu Ocean University, Lianyungang 222005, China
2. Co-Innovation Center of Jiangsu Marine Bio-industry Technology, Jiangsu Ocean University, Lianyungang 222005, China
-
Received:2023-02-28Revised:2023-05-16Online:2023-06-20Published:2023-05-29 -
Contact:Xing Chaofan E-mail:chaofanxing2021@163.com -
Supported by:National Natural Science Foundation of China(32200411);Natural Science Foundation of the Jiangsu Higher Education Institutions(22KJB180014);Priority Academic Program Development Fund of Jiangsu Higher Education Institutions(22KJB180014)
Cite this article
Chaofan Xing, Mintao Wang, Lei Wang, Xin Shen. Progress on the mechanism of left-right asymmetrical patterning in bilaterians[J]. Hereditas(Beijing), 2023, 45(6): 488-500.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
Table 1
Establishment of left-right asymmetrical signal in several vertebrates and invertebrates"
| 分类 | 物种 | Dand5 | Nodal | Vg1 | Lefty | Pitx | BMP信号 |
|---|---|---|---|---|---|---|---|
| 脊椎动物 | 小鼠 (Mus musculus) | 参与 | 参与 | 参与 | 参与 | 参与 | 参与 |
| 鸡 (Gallus gallus) | 参与 | 参与 | 参与 | 参与 | 参与 | 参与 | |
| 非洲爪蟾 (Xenopus laevis) | 参与 | 参与 | 参与 | 参与 | 参与 | 参与 | |
| 斑马鱼 (Danio rerio) | 参与 | 参与 | 参与 | 参与 | 不参与 | 参与 | |
| 无脊椎动物 | 住囊虫 (Oikopleura) | 未知 | 不存在 | 未知 | 未知 | 不参与 | 参与 |
| 海鞘 (Ascidiacea) | 未知 | 参与 | 未知 | 未知 | 参与 | 未知 | |
| 文昌鱼 (Branchiostoma) | 参与 | 参与 | 参与 | 参与 | 参与 | 参与 | |
| 海胆 (Echinoidea) | 未知 | 参与 | 未知 | 参与 | 参与 | 参与 | |
| 果蝇 (Drosophila) | 未知 | 不存在 | 未知 | 未知 | 不存在 | 未知 | |
| 腹足类 (Snails) | 未知 | 参与 | 未知 | 未知 | 参与 | 参与 | |
| 线虫 (Nematoda) | 未知 | 不存在 | 未知 | 未知 | 不存在 | 未知 | |
| 水螅 (Hydra) | 未知 | 参与 | 未知 | 未知 | 参与 | 未知 |
| [1] | Rebecca DB, Caspary T. Left-right asymmetry: lessons from Cancún. Development, 2013, 140(22): 4465-4470. |
| [2] |
Namigai EKO, Kenny NJ, Shimeld SM. Right across the tree of life: the evolution of left-right asymmetry in the Bilateria. Genesis, 2014, 52(6): 458-470.
doi: 10.1002/dvg.22748 pmid: 24510729 |
| [3] |
Kuroda R, Endo B, Abe M, Shimizu M. Chiral blastomere arrangement dictates zygotic left-right asymmetry pathway in snails. Nature, 2009, 462(7274): 790-794.
doi: 10.1038/nature08597 |
| [4] |
Spéder P, Ádám G, Noselli S. Type ID unconventional myosin controls left-right asymmetry in Drosophila. Nature, 2006, 440(7085): 803-807.
doi: 10.1038/nature04623 |
| [5] |
Eaves AA. Potential for paired vestibules in plutei (Echinodermata, Echinoidea). Invertebrate Biology, 2005, 124(2): 174-184.
doi: 10.1111/j.1744-7410.2005.00017.x |
| [6] | Smith MM, Cruz Smith L, Cameron RA, Urry LA. The larval stages of the sea urchin, Strongylocentrotus purpuratus. J Morphol, 2008, 269(6): 713-733. |
| [7] |
Hu GW, Zhang ZZ, Gao H. Progress on the left-right asymmetry patterning in amphioxus. Hereditas(Beijing), 2021, 43(2): 134-141.
doi: 10.16288/j.yczz.20-246 pmid: 33724216 |
|
胡广伟, 张珍珍, 高焕. 文昌鱼左右体轴建立机制的研究进展. 遗传, 2021, 43(2): 134-141.
doi: 10.16288/j.yczz.20-246 pmid: 33724216 |
|
| [8] |
Soukup V, Yong LW, Lu TM, Huang SW, Kozmik Z, Yu JK. The Nodal signaling pathway controls left-right asymmetric development in amphioxus. EvoDevo, 2015, 6(1): 1-23.
doi: 10.1186/2041-9139-6-1 |
| [9] |
Hamada H, Meno C, Watanabe D, Saijoh Y. Establishment of vertebrate left-right asymmetry. Nat Rev Genet, 2002, 3(2): 103-113.
doi: 10.1038/nrg732 pmid: 11836504 |
| [10] |
Blum M, Feistel K, Thumberger T, Schweickert A. The evolution and conservation of left-right patterning mechanisms. Development, 2014, 141(8): 1603-1613.
doi: 10.1242/dev.100560 pmid: 24715452 |
| [11] |
Blum M, Ott T. Animal left-right asymmetry. Curr Biol, 2018, 28(7): R301-R304.
doi: 10.1016/j.cub.2018.02.073 |
| [12] |
Isao S, Hajime I. Human heterotaxy syndrome—from molecular genetics to clinical features, management, and prognosis. Circ J, 2012, 76(9): 2066-2075.
doi: 10.1253/circj.CJ-12-0957 |
| [13] |
Kennedy MP, Omran H, Leigh MW, Dell S, Morgan L, Molina PL, Robinson BV, Minnix SL, Olbrich H, Severin T, Ahrens P, Lange L, Morillas HN, Noone PG, Zariwala MA, Knowles MR. Congenital heart disease and other heterotaxic defects in a large cohort of patients with primary ciliary dyskinesia. Circulation, 2007, 115(22): 2814-2821.
doi: 10.1161/CIRCULATIONAHA.106.649038 pmid: 17515466 |
| [14] |
Nakamura T, Hamada H. Left-right patterning: conserved and divergent mechanisms. Development, 2012, 139(18): 3257-3262.
doi: 10.1242/dev.061606 pmid: 22912409 |
| [15] | Hamada H, Tam P.Diversity of left-right symmetry breaking strategy in animals. F1000Res, 2020, 9:F1000 Faculty Rev-123. |
| [16] | Zhang YJ, Xu PF. Research progress of left-right asymmetry development in vertebrates. J Zhejiang Univ Agric Life Sci, 2020, 46(6): 647-659. |
| 张颖洁, 徐鹏飞. 脊椎动物左右不对称发育研究进展. 浙江大学学报, 2020, 46(6): 647-659. | |
| [17] |
Raya A, Belmonte JCI. Left-right asymmetry in the vertebrate embryo: from early information to higher-level integration. Nat Rev Genet, 2006, 7(4): 283-293.
pmid: 16543932 |
| [18] |
Hirokawa N, Tanaka Y, Okada Y, Takeda S. Nodal flow and the generation of left-right asymmetry. Cell, 2006, 125(1): 33-45.
doi: 10.1016/j.cell.2006.03.002 pmid: 16615888 |
| [19] |
Schweickert A, Vick P, Getwan M, Weber T, Schneider I, Eberhardt M, Beyer T, Pachur A, Blum M. The Nodal inhibitor Coco is a critical target of leftward flow in Xenopus. Curr Biol, 2010, 20(8): 738-743.
doi: 10.1016/j.cub.2010.02.061 pmid: 20381352 |
| [20] |
Gros J, Feistel K, Viebahn C, Blum M, Tabin CJ. Cell movements at Hensen’s node establish left/right asymmetric gene expression in the chick. Science, 2009, 324(5929): 941-944.
doi: 10.1126/science.1172478 |
| [21] |
Grimes DT, Burdine RD. Left-right patterning: breaking symmetry to asymmetric morphogenesis. Trends Genet, 2017, 33(9): 616-628.
doi: S0168-9525(17)30102-6 pmid: 28720483 |
| [22] |
Amack JD. Salient features of the ciliated organ of asymmetry. Bioarchitecture, 2014, 4(1): 6-15.
doi: 10.4161/bioa.28014 pmid: 24481178 |
| [23] |
Daems M, Peacock HM, Jones EA. Fluid flow as a driver of embryonic morphogenesis. Development, 2020, 147(15): dev185579.
doi: 10.1242/dev.185579 |
| [24] |
McGrath J, Somlo S, Makova S, Tian X, Brueckner M. Two populations of node monocilia initiate left-right asymmetry in the mouse. Cell, 2003, 114(1): 61-73.
pmid: 12859898 |
| [25] |
Nonaka S, Shiratori H, Saijoh Y, Hamada H. Determination of left-right patterning of the mouse embryo by artificial nodal flow. Nature, 2002, 418(6893): 96-99.
doi: 10.1038/nature00849 |
| [26] |
Okabe N, Xu B, Burdine RD. Fluid dynamics in zebrafish Kupffer's vesicle. Dev Dyn, 2008, 237(12): 3602-3612.
doi: 10.1002/dvdy.v237:12 |
| [27] |
Hashimoto M, Shinohara K, Wang JB, Ikeuchi S, Yoshiba S, Meno C, Nonaka S, Takada S, Hatta K, Wynshaw-Boris A. Planar polarization of node cells determines the rotational axis of node cilia. Nat Cell Biol, 2010, 12(2): 170-176.
doi: 10.1038/ncb2020 pmid: 20098415 |
| [28] |
Antic D, Stubbs JL, Suyama K, Kintner C, Scott MP, Axelrod JD. Planar cell polarity enables posterior localization of nodal cilia and left-right axis determination during mouse and Xenopus embryogenesis. PloS One, 2010, 5(2): e8999.
doi: 10.1371/journal.pone.0008999 |
| [29] |
Song H, Hu JX, Chen W, Elliott G, Andre P, Gao B, Yang YZ. Planar cell polarity breaks bilateral symmetry by controlling ciliary positioning. Nature, 2010, 466(7304): 378-382.
doi: 10.1038/nature09129 |
| [30] |
Borovina A, Superina S, Voskas D, Ciruna B. Vangl2 directs the posterior tilting and asymmetric localization of motile primary cilia. Nat Cell Biol, 2010, 12(4):407-412.
doi: 10.1038/ncb2042 pmid: 20305649 |
| [31] | Minegishi K, Hashimoto M, Ajima R, Takaoka K, Shinohara K, Ikawa Y, Nishimura H, Mcmahon AP, Willert K, Okada Y, Sasaki H, Shi DB, Fujimori T, Ohtsuka T, Igarashi Y, Yamaguchi TP, Shimono A, Shiratori H, Hamada H. A Wnt5 activity asymmetry and intercellular signaling via PCP proteins polarize node cells for left-right symmetry breaking. Dev Cell, 2017, 40(5): 439-452.e4. |
| [32] |
Katoh TA, Omori T, Mizuno K, Sai XR, Minegishi K, Ikawa Y, Nishimura H, Itabashi T, Kajikawa E, Hiver S, Iwane AH, Ishikawa T, Okada Y, Nishizaka T, Hamada H. Immotile cilia mechanically sense the direction of fluid flow for left-right determination. Science, 2023, 379(6627): 66-71.
doi: 10.1126/science.abq8148 pmid: 36603091 |
| [33] |
Djenoune L, Mahamdeh M, Truong TV, Nguyen CT, Fraser SE, Brueckner M, Howard J, Yuan S. Cilia function as calcium-mediated mechanosensors that instruct left-right asymmetry. Science, 2023, 379(6627): 71-78.
doi: 10.1126/science.abq7317 pmid: 36603098 |
| [34] |
Tavares B, Jacinto R, Sampaio P, Pestana S, Pinto A, Vaz A, Roxo-Rosa M, Gardner R, Lopes T, Schilling B, Henry I, Saúde L, Lopes SS. Notch/Her12 signalling modulates, motile/immotile cilia ratio downstream of Foxj1a in zebrafish left-right organizer. eLife, 2017, 6: e25165.
doi: 10.7554/eLife.25165 |
| [35] |
Katsu K, Tokumori D, Tatsumi N, Suzuki A, Yokouchi Y. BMP inhibition by DAN in Hensen's node is a critical step for the establishment of left-right asymmetry in the chick embryo. Dev Biol, 2012, 363(1): 15-26.
doi: 10.1016/j.ydbio.2011.12.015 pmid: 22202776 |
| [36] |
Kajikawa E, Horo U, Ide T, Mizuno K, Minegishi K, Hara Y, Ikawa Y, Nishimura H, Uchikawa M, Kiyonari H, Kuraku S, Hamada H. Nodal paralogues underlie distinct mechanisms for visceral left-right asymmetry in reptiles and mammals. Nat Ecol Evol, 2020, 4(2): 261-269.
doi: 10.1038/s41559-019-1072-2 pmid: 31907383 |
| [37] |
Nishide K, Mugitani M, Kumano G, Nishida H. Neurula rotation determines left-right asymmetry in ascidian tadpole larvae. Development, 2012, 139(8): 1467-1475.
doi: 10.1242/dev.076083 pmid: 22399684 |
| [38] |
Yamada S, Tanaka Y, Imai KS, Saigou M, Onuma TA, Nishida H. Wavy movements of epidermis monocilia drive the neurula rotation that determines left-right asymmetry in ascidian embryos. Dev Biol, 2019, 448(2): 173-182.
doi: S0012-1606(18)30215-X pmid: 30059669 |
| [39] |
Li G, Liu X, Xing CF, Zhang HY, Shimeld SM, Wang YQ. Cerberus-Nodal-Lefty-Pitx signaling cascade controls left-right asymmetry in amphioxus. Proc Natl Acad Sci USA, 2017, 114(14): 3684-3689.
doi: 10.1073/pnas.1620519114 |
| [40] |
Wang H, Li G, Wang YQ. Generating amphioxus Hedgehog knockout mutants and phenotype analysis. Hereditas(Beijing), 2015, 37(10):1036-1043.
doi: 10.16288/j.yczz.15-276 pmid: 26496756 |
|
王慧, 李光, 王义权. 文昌鱼Hedgehog基因敲除和突变体表型分析. 遗传, 2015, 37(10):1036-1043.
doi: 10.16288/j.yczz.15-276 pmid: 26496756 |
|
| [41] | Zhu X, Shi CG, Zhong YH, Liu X, Yan QN, Wu XT, Wang YQ, Li G. Cilia-driven asymmetric Hedgehog signalling determines the amphioxus left-right axis by controlling Dand5expression. Development, 2020, 147(1): dev182469. |
| [42] |
Tisler M, Wetzel F, Mantino S, Kremnyov S, Thumberger T, Schweickert A, Blum M, Vick P. Cilia are required for asymmetric nodal induction in the sea urchin embryo. BMC Dev Biol, 2016, 16(1): 28.
doi: 10.1186/s12861-016-0128-7 pmid: 27553781 |
| [43] |
Kuroda R, Fujikura K, Abe M, Hosoiri Y, Asakawa S, Shimizu M, Umeda S, Ichikawa F, Takahashi H. Diaphanous gene mutation affects spiral cleavage and chirality in snails. Sci Rep, 2016, 6(1): 34809.
doi: 10.1038/srep34809 |
| [44] |
Abe M, Kuroda R. The development of CRISPR for a mollusc establishes the formin Lsdia1 as the long-sought gene for snail dextral/sinistral coiling. Development, 2019, 146(9): dev175976.
doi: 10.1242/dev.175976 |
| [45] |
González-Morales N, Géminard C, Lebreton G, Cerezo D, Coutelis J-B, Noselli S. The atypical cadherin dachsous controls left-right asymmetry in Drosophila. Dev Cell, 2015, 33(6): 675-689.
doi: 10.1016/j.devcel.2015.04.026 pmid: 26073018 |
| [46] |
Lebreton G, Géminard C, Lapraz F, Pyrpassopoulos S, Cerezo D, Spéder P, Ostap EM, Noselli S. Molecular to organismal chirality is induced by the conserved myosin 1D. Science, 2018, 362(6417): 949-952.
doi: 10.1126/science.aat8642 pmid: 30467170 |
| [47] |
Pohl C, Bao ZR. Chiral forces organize left-right patterning in C. elegans by uncoupling midline and anteroposterior axis. Dev Cell, 2010, 19(3): 402-412.
doi: 10.1016/j.devcel.2010.08.014 |
| [48] |
Pohl C. Left-right patterning in the C. elegans embryo: unique mechanisms and common principles. Commun Integr Biol, 2011, 4(1): 34-40.
doi: 10.4161/cib.14144 |
| [49] |
Naganathan SR, Fürthauer S, Nishikawa M, Jülicher F, Grill SW. Active torque generation by the actomyosin cell cortex drives left-right symmetry breaking. eLife, 2014, 3: e04165.
doi: 10.7554/eLife.04165 |
| [50] |
Yuan S, Brueckner M. Left-right asymmetry: myosin 1D at the center. Curr Biol, 2018, 28(9): R567-R569.
doi: 10.1016/j.cub.2018.03.019 |
| [51] |
Rankin CT, Bunton T, Lawler AM, Lee SJ.Regulation of left-right patterning in mice by growth/differentiation factor-1. Nat Genet, 2000, 24(3): 262-265.
pmid: 10700179 |
| [52] |
Pelliccia JL, Jindal GA, Burdine RD. Gdf3 is required for robust Nodal signaling during germ layer formation and left-right patterning. eLife, 2017, 6: e28635.
doi: 10.7554/eLife.28635 |
| [53] |
Marques S, Borges AC, Silva AC, Freitas S, Cordenonsi M, Belo JA. The activity of the Nodal antagonist Cerl-2 in the mouse node is required for correct L/R body axis. Genes Dev, 2004, 18(19): 2342-2347.
doi: 10.1101/gad.306504 |
| [54] |
Tavares AT, Andrade S, Silva AC, Belo JA. Cerberus is a feedback inhibitor of Nodal asymmetric signaling in the chick embryo. Development, 2007, 134(11): 2051- 2060.
pmid: 17507406 |
| [55] |
Meno C, Shimono A, Saijoh Y, Yashiro K, Mochida K, Ohishi S, Noji S, Kondoh H, Hamada H. Lefty-1 is required for left-right determination as a regulator of lefty-2 and nodal. Cell, 1998, 94(3): 287-297.
pmid: 9708731 |
| [56] |
Wang XH, Yost HJ. Initiation and propagation of posterior to anterior (PA) waves in zebrafish left-right development. Dev Dyn, 2008, 237(12): 3640-3647.
doi: 10.1002/dvdy.v237:12 |
| [57] |
Smith KA, Noël E, Thurlings I, Rehmann H, Chocron S, Bakkers J. Bmp and nodal independently regulate lefty1 expression to maintain unilateral nodal activity during left-right axis specification in zebrafish. PLoS Genet, 2011, 7(9): e1002289.
doi: 10.1371/journal.pgen.1002289 |
| [58] |
Shiratori H, Sakuma R, Watanabe M, Hashiguchi H, Mochida K, Sakai Y, Nishino J, Saijoh Y, Whitman M, Hamada H. Two-step regulation of left-right asymmetric expression of Pitx2: initiation by nodal signaling and maintenance by Nkx2. Mol Cell, 2001, 7(1): 137-149.
pmid: 11172719 |
| [59] |
Shiratori H, Yashiro K, Shen MM, Hamada H. Conserved regulation and role of Pitx2 in situs-specific morphogenesis of visceral organs. Development, 2006, 133(15): 3015-3025.
pmid: 16835440 |
| [60] |
Morokuma J, Ueno M, Kawanishi H, Saiga H, Nishida H. HrNodal, the ascidian nodal-related gene, is expressed in the left side of the epidermis, and lies upstream of HrPitx. Dev Genes Evol, 2002, 212(9): 439-446.
pmid: 12373589 |
| [61] |
Yoshida K, Saiga H. Left-right asymmetric expression of Pitx is regulated by the asymmetric Nodal signaling through an intronic enhancer in Ciona intestinalis. Dev Genes Evol, 2008, 218(7): 353-360.
doi: 10.1007/s00427-008-0230-3 pmid: 18546017 |
| [62] |
Soukup V, Yong LW, Lu TM, Huang SW, Kozmik Z, Yu JK. The Nodal signaling pathway controls left-right asymmetric development in amphioxus. EvoDevo, 2015, 6(1): 5.
doi: 10.1186/2041-9139-6-5 |
| [63] |
Xing CF, Pan RR, Hu GW, Liu X, Wang YQ, Li G. Pitx controls amphioxus asymmetric morphogenesis by promoting left-side development and repressing right-side formation. BMC Biol, 2021, 19(1): 166.
doi: 10.1186/s12915-021-01095-0 |
| [64] |
Duboc V, Röttinger E, Lapraz F, Besnardeau L, Lepage T. Left-right asymmetry in the sea urchin embryo is regulated by nodal signaling on the right side. Dev Cell, 2005, 9(1): 147-158.
pmid: 15992548 |
| [65] |
Molina MD, De Crozé N, Haillot E, Lepage T. Nodal: master and commander of the dorsal-ventral and left-right axes in the sea urchin embryo. Curr Opin Genet Dev, 2013, 23(4): 445-453.
doi: 10.1016/j.gde.2013.04.010 pmid: 23769944 |
| [66] |
Su YH. Telling left from right: left-right asymmetric controls in sea urchins. Genesis, 2014, 52(3): 269-278.
doi: 10.1002/dvg.22739 |
| [67] |
Range R, Lapraz F, Quirin M, Marro S, Besnardeau L, Lepage T.Cis-regulatory analysis of nodal and maternal control of dorsal-ventral axis formation by Univin, a TGF-β related to Vg1. Development, 2007, 134(20): 3649-3664.
pmid: 17855430 |
| [68] |
Range R, Lepage T. Maternal Oct1/2 is required for Nodal and Vg1/Univin expression during dorsal-ventral axis specification in the sea urchin embryo. Dev Biol, 2011, 357(2): 440-449.
doi: 10.1016/j.ydbio.2011.07.005 pmid: 21782809 |
| [69] |
Grande C, Patel NH. Nodal signalling is involved in left-right asymmetry in snails. Nature, 2009, 457(7232): 1007-1011.
doi: 10.1038/nature07603 |
| [70] |
Watanabe H, Schmidt HA, Kuhn A, Höger SK, Kocagöz Y, Laumann-Lipp N, Ozbek S, Holstein TW. Nodal signalling determines biradial asymmetry in Hydra. Nature, 2014, 515(7525): 112-115.
doi: 10.1038/nature13666 |
| [71] |
Cox CJ, Espinoza HM, Mcwilliams B, Chappell K, Morton L, Hjalt TA, Semina EV, Amendt BA. Differential regulation of gene expression by PITX2 isoforms. J Biol Chem, 2002, 277(28): 25001-25010.
doi: 10.1074/jbc.M201737200 pmid: 11948188 |
| [72] |
Schweickert A, Campione M, Steinbeisser H, Blum M. Pitx2 isoforms: involvement of Pitx2c but not Pitx2a or Pitx2b in vertebrate left-right asymmetry. Mech Dev, 2000, 90(1): 41-51.
doi: 10.1016/S0925-4773(99)00227-0 |
| [73] |
Yu X, St Amand TR, Wang S, Li G, Zhang Y, Hu Y, Nguyen L, Qiu MS, Chen YP. Differential expression and functional analysis of Pitx2 isoforms in regulation of heart looping in the chick. Development, 2001, 128(6): 1005-1013.
doi: 10.1242/dev.128.6.1005 pmid: 11222154 |
| [74] |
Essner JJ, Branford WW, Zhang J, Yost HJ. Mesendoderm and left-right brain, heart and gut development are differentially regulated by Pitx2 isoforms. Development, 2000, 127(5): 1081-1093.
doi: 10.1242/dev.127.5.1081 pmid: 10662647 |
| [75] |
Yoshioka H, Meno C, Koshiba K, Sugihara M, Itoh H, Ishimaru Y, Inoue T, Ohuchi H, Semina EV, Murray JC, Hamada H, Noji S. Pitx2, a bicoid-type homeobox gene, is involved in a lefty-signaling pathway in determination of left-right asymmetry. Cell, 1998, 94(3): 299-305.
pmid: 9708732 |
| [76] |
Lin CR, Kioussi C, O'connell S, Briata P, Szeto D, Liu F, Izpisúa-Belmonte JC, Rosenfeld MG. Pitx2 regulates lung asymmetry, cardiac positioning and pituitary and tooth morphogenesis. Nature, 1999, 401(6750): 279-282.
doi: 10.1038/45803 |
| [77] |
Davis NM, Kurpios NA, Sun XX, Gros J, Martin JF, Tabin CJ. The chirality of gut rotation derives from left-right asymmetric changes in the architecture of the dorsal mesentery. Dev Cell, 2008, 15(1): 134-145.
doi: 10.1016/j.devcel.2008.05.001 pmid: 18606147 |
| [78] |
Davis A, Amin NM, Johnson C, Bagley K, Ghashghaei HT, Nascone-Yoder N. Stomach curvature is generated by left-right asymmetric gut morphogenesis. Development, 2017, 144(8): 1477-1483.
doi: 10.1242/dev.143701 pmid: 28242610 |
| [79] |
Ryan AK, Blumberg B, Rodriguez-Esteban C, Yonei- Tamura S, Tamura K, Tsukui T, De La Peña J, Sabbagh WS, Greenwald J, Choe S, Norris DP, Robertson EJ, Evans RM, Rosenfeld MG, Izpisúa Belmonte JC. Pitx2 determines left-right asymmetry of internal organs in vertebrates. Nature, 1998, 394(6693): 545-551.
doi: 10.1038/29004 |
| [80] |
Womble M, Amin NM, Nascone-Yoder N. The left-right asymmetry of liver lobation is generated by Pitx2c- mediated asymmetries in the hepatic diverticulum. Dev Biol, 2018, 439(2): 80-91.
doi: S0012-1606(17)30675-9 pmid: 29709601 |
| [81] |
Liu Y, Semina EV. Pitx2 deficiency results in abnormal ocular and craniofacial development in zebrafish. PLoS One, 2012, 7(1): e30896.
doi: 10.1371/journal.pone.0030896 |
| [82] |
Ji YC, Buel SM, Amack JD. Mutations in zebrafish pitx2 model congenital malformations in Axenfeld-Rieger syndrome but do not disrupt left-right placement of visceral organs. Dev Biol, 2016, 416(1): 69-81.
doi: 10.1016/j.ydbio.2016.06.010 |
| [83] |
Welsh IC, Thomsen M, Gludish DW, Alfonso-Parra C, Bai Y, Martin JF, Kurpios NA. Integration of left-right Pitx2 transcription and Wnt signaling drives asymmetric gut morphogenesis via Daam2. Dev Cell, 2013, 26(6): 629-644.
doi: 10.1016/j.devcel.2013.07.019 pmid: 24091014 |
| [84] |
Sanketi BD, Zuela-Sopilniak N, Bundschuh E, Gopal S, Hu S, Long J, Lammerding J, Hopyan S, Kurpios NA. Pitx2 patterns an accelerator-brake mechanical feedback through latent TGFβ to rotate the gut. Science, 2022, 377(6613): eabl3921.
doi: 10.1126/science.abl3921 |
| [85] |
Martin JF. Left right asymmetry, the pulmonary vein, and a-fib. Circ Res, 2007, 101(9): 853-855.
pmid: 17967793 |
| [86] |
Wang J, Klysik E, Sood S, Johnson RL, Wehrens XHT, Martin JF. Pitx2 prevents susceptibility to atrial arrhythmias by inhibiting left-sided pacemaker specification. Proc Natl Acad Sci USA, 2010, 107(21): 9753-9758.
doi: 10.1073/pnas.0912585107 pmid: 20457925 |
| [87] |
Shen MM. Nodal signaling: developmental roles and regulation. Development, 2007, 134(6): 1023-1034.
doi: 10.1242/dev.000166 pmid: 17287255 |
| [88] |
Zinski J, Tajer B, Mullins MC. TGF-β family signaling in early vertebrate development. Cold Spring Harb Perspect Biol, 2018, 10(6): a033274.
doi: 10.1101/cshperspect.a033274 |
| [89] |
Brennan J, Norris DP, Robertson E. Nodal activity in the node governs left-right asymmetry. Genes Dev, 2002, 16(18): 2339-2344.
doi: 10.1101/gad.1016202 |
| [90] |
Long S, Ahmad N, Rebagliati M. The zebrafish nodal-related gene southpaw is required for visceral and diencephalic left-right asymmetry. Development, 2003, 130(11): 2303-2316.
doi: 10.1242/dev.00436 |
| [91] |
Noël ES, Verhoeven M, Lagendijk AK, Tessadori F, Smith K, Choorapoikayil S, Den Hertog J, Bakkers J. A Nodal-independent and tissue-intrinsic mechanism controls heart-looping chirality. Nat Commun, 2013, 4(1): 2754.
doi: 10.1038/ncomms3754 |
| [92] |
Rago L, Castroviejo N, Fazilaty H, Garcia-Asencio F, Nieto MA, Ocaña OH, Galcerán J. MicroRNAs establish the right-handed dominance of the heart laterality pathway in vertebrates. Dev Cell, 2019, 51(4): 446-459.
doi: S1534-5807(19)30768-3 pmid: 31630980 |
| [93] |
Roussigne M, Blader P, Wilson SW. Breaking symmetry: the zebrafish as a model for understanding left-right asymmetry in the developing brain. Dev Neurobiol, 2012, 72(3): 269-281.
pmid: 22553774 |
| [94] |
Mine N, Anderson RM, Klingensmith J. BMP antagonism is required in both the node and lateral plate mesoderm for mammalian left-right axis establishment. Development, 2008, 135(14): 2425-2434.
doi: 10.1242/dev.018986 pmid: 18550712 |
| [95] |
Capdevila J, Vogan KJ, Tabin CJ, Izpisúa Belmonte JC. Mechanisms of left-right determination in vertebrates. Cell, 2000, 101(1): 9-21.
pmid: 10778851 |
| [96] |
Ocaña OH, Coskun H, Minguillón C, Murawala P, Tanaka EM, Galcerán J, Muñoz-Chápuli R, Nieto MA. A right-handed signalling pathway drives heart looping in vertebrates. Nature, 2017, 549(7670): 86-90.
doi: 10.1038/nature23454 |
| [97] |
Tessadori F, Barske L, Nelson N, Algra HA, Willekers S, Nichols JT, Crump JG, Bakkers J. Zebrafish prrx1a mutants have normal hearts. Nature, 2020, 585(7826): E14-E16.
doi: 10.1038/s41586-020-2674-1 |
| [98] |
Onuma TA, Hayashi M, Gyoja F, Kishi K, Wang K, Nishida H. A chordate species lacking Nodal utilizes calcium oscillation and Bmp for left-right patterning. Proc Natl Acad Sci USA, 2020, 117(8): 4188-4198.
doi: 10.1073/pnas.1916858117 |
| [99] |
Soukup V, Kozmik Z. The Bmp signaling pathway regulates development of left-right asymmetry in amphioxus. Dev Biol, 2018, 434(1): 164-174.
doi: S0012-1606(17)30153-7 pmid: 29224891 |
| [100] |
Luo YJ, Su YH. Opposing Nodal and BMP signals regulate left-right asymmetry in the sea urchin larva. PLoS Biol, 2012, 10(10): e1001402.
doi: 10.1371/journal.pbio.1001402 |
| [101] |
Chougule A, Lapraz F, Földi I, Cerezo D, Mihály J, Noselli S. The Drosophila actin nucleator DAAM is essential for left-right asymmetry. PLoS Genet, 2020, 16(4): e1008758.
doi: 10.1371/journal.pgen.1008758 |
| [102] |
Shimizu K, Sarashina I, Kagi H, Endo K. Possible functions of Dpp in gastropod shell formation and shell coiling. Dev Genes Evol, 2011, 221(2): 59-68.
doi: 10.1007/s00427-011-0358-4 pmid: 21556857 |
| [1] | Shan He, Jian Zhao, Xiaofeng Song. Effects of N6-methyladenosine modification on the function of the female reproductive system [J]. Hereditas(Beijing), 2023, 45(6): 472-487. |
| [2] | Feifei Li, Yanmin Hao, Minlong Cui, Chunlan Piao. Cloning and functional analysis of RADIALIS-like 1 gene from Antirrhinum majus [J]. Hereditas(Beijing), 2023, 45(6): 526-535. |
| [3] | Mengxuan Xu, Ming Zhou. Advances of RNA polymerase IV in controlling DNA methylation and development in plants [J]. Hereditas(Beijing), 2022, 44(7): 567-580. |
| [4] | Sinan Luan, Lele Liu, Jiayuan Zhou, Nurasya Imam, Minlong Cui, Chunlan Piao. Functional analysis of flower development related gene SvGLOBOSA from Senecio vulgaris [J]. Hereditas(Beijing), 2022, 44(6): 521-530. |
| [5] | Yuan Zhang, Yuting Zhao, Lenan Zhuang, Jin He. Transcriptional regulation of transcriptional Mediator complexes in cardiovascular development and disease [J]. Hereditas(Beijing), 2022, 44(5): 383-397. |
| [6] | Mengxiao Wang, Margaret S. Ho. Recent advances on the role of glia in physiological behaviors: insights from Drosophila melanogaster [J]. Hereditas(Beijing), 2022, 44(4): 300-312. |
| [7] | Hui Qu, Yi Liu, Yawen Chen, Hui Wang. Alteration of imprinted genes and offspring organ development caused by environmental factors [J]. Hereditas(Beijing), 2022, 44(2): 107-116. |
| [8] | Ke Mao, Ziqiu Meng, Yongbiao Zhang. Progress on the regulation of neural crest and the genetics in craniofacial development [J]. Hereditas(Beijing), 2022, 44(12): 1089-1102. |
| [9] | Liwen Zhang, Meihua Ruan, Jialan Liu, Caihong He, Jianrong Yu. Progress on research and development in diabetes mellitus [J]. Hereditas(Beijing), 2022, 44(10): 824-839. |
| [10] | Zhijing Zhang, Yu Qiao, Yuchen Sun, Lei Lei. Epigenetic “reader” BET proteins regulate mammalian development and iPSC reprogramming [J]. Hereditas(Beijing), 2022, 44(1): 36-45. |
| [11] | Yan Zhu, Ming Wei, Xiao Zhou, Linhua Deng, Jian Qiu, Guo Li, Shiqiang Zhou, Hao Xie, Desheng Li, Chengdong Wang. Progress on miRNA in giant panda (Ailuropoda melanoleuca) [J]. Hereditas(Beijing), 2021, 43(9): 849-857. |
| [12] | Caihong He, Wanzi Jiang, Liwen Zhang, Meihua Ruan, Hongwen Zhou, Jianrong Yu. Current status and future perspectives of rare disease research [J]. Hereditas(Beijing), 2021, 43(6): 531-544. |
| [13] | Wei Xie, Chengqi Lin, Jian Li, Moyi Li. Decoding brain encoded by genome [J]. Hereditas(Beijing), 2021, 43(5): 393-396. |
| [14] | Wenjing Zhu, Zhiwei Liu. MDMPR: Mouse Developmental and Metabolic Phenotype Repository [J]. Hereditas(Beijing), 2021, 43(4): 375-384. |
| [15] |
Guangwei Hu, Zhenzhen Zhang, Huan Gao.
|
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||